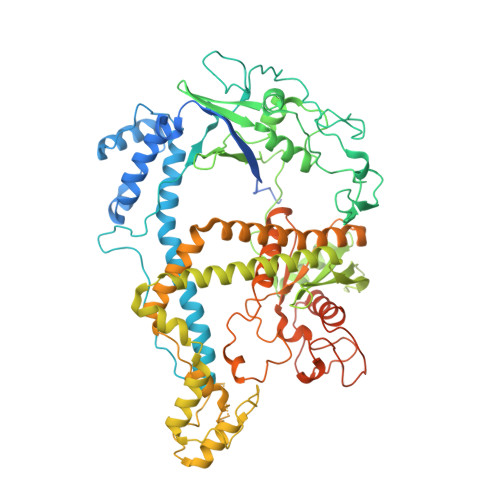
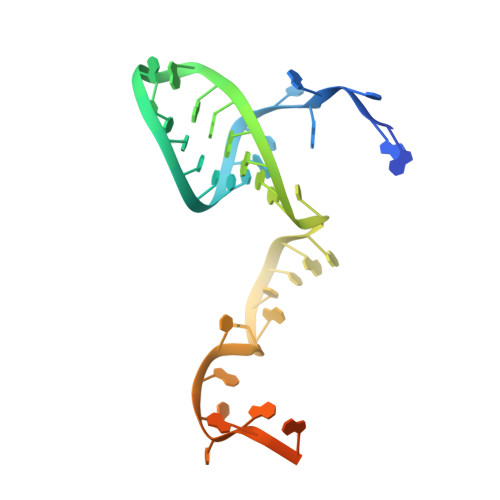
DNA interference states of the hypercompact CRISPR-Cas Phi effector.
Pausch, P., Soczek, K.M., Herbst, D.A., Tsuchida, C.A., Al-Shayeb, B., Banfield, J.F., Nogales, E., Doudna, J.A.(2021) Nat Struct Mol Biol 28: 652-661
- PubMed: 34381246 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-021-00632-3
- Primary Citation Related Structures:
7LYS, 7LYT, 7M5O - PubMed Abstract:
CRISPR-CasΦ, a small RNA-guided enzyme found uniquely in bacteriophages, achieves programmable DNA cutting as well as genome editing. To investigate how the hypercompact enzyme recognizes and cleaves double-stranded DNA, we determined cryo-EM structures of CasΦ (Cas12j) in pre- and post-DNA-binding states. The structures reveal a streamlined protein architecture that tightly encircles the CRISPR RNA and DNA target to capture, unwind and cleave DNA. Comparison of the pre- and post-DNA-binding states reveals how the protein rearranges for DNA cleavage upon target recognition. On the basis of these structures, we created and tested mutant forms of CasΦ that cut DNA up to 20-fold faster relative to wild type, showing how this system may be naturally attenuated to improve the fidelity of DNA interference. The structural and mechanistic insights into how CasΦ binds and cleaves DNA should allow for protein engineering for both in vitro diagnostics and genome editing.
- Innovative Genomics Institute, University of California, Berkeley, Berkeley, CA, USA.
Organizational Affiliation: