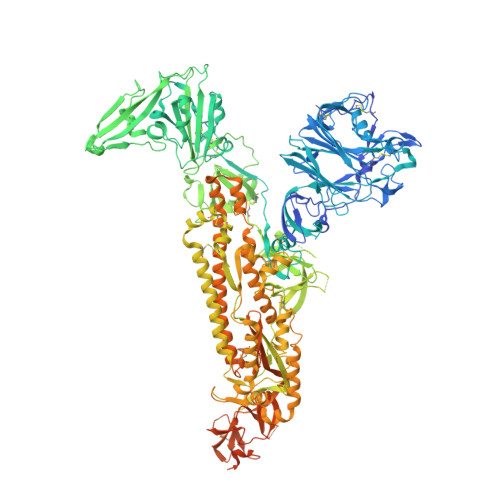
Structural basis for broad coronavirus neutralization.
Sauer, M.M., Tortorici, M.A., Park, Y.J., Walls, A.C., Homad, L., Acton, O.J., Bowen, J.E., Wang, C., Xiong, X., de van der Schueren, W., Quispe, J., Hoffstrom, B.G., Bosch, B.J., McGuire, A.T., Veesler, D.(2021) Nat Struct Mol Biol 28: 478-486
- PubMed: 33981021 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-021-00596-4
- Primary Citation Related Structures:
7M51, 7M52, 7M53, 7M55, 7M5E - PubMed Abstract:
Three highly pathogenic β-coronaviruses have crossed the animal-to-human species barrier in the past two decades: SARS-CoV, MERS-CoV and SARS-CoV-2. To evaluate the possibility of identifying antibodies with broad neutralizing activity, we isolated a monoclonal antibody, termed B6, that cross-reacts with eight β-coronavirus spike glycoproteins, including all five human-infecting β-coronaviruses. B6 broadly neutralizes entry of pseudotyped viruses from lineages A and C, but not from lineage B, and the latter includes SARS-CoV and SARS-CoV-2. Cryo-EM, X-ray crystallography and membrane fusion assays reveal that B6 binds to a conserved cryptic epitope located in the fusion machinery. The data indicate that antibody binding sterically interferes with the spike conformational changes leading to membrane fusion. Our data provide a structural framework explaining B6 cross-reactivity with β-coronaviruses from three lineages, along with a proof of concept for antibody-mediated broad coronavirus neutralization elicited through vaccination. This study unveils an unexpected target for next-generation structure-guided design of a pan-β-coronavirus vaccine.
- Department of Biochemistry, University of Washington, Seattle, WA, USA.
Organizational Affiliation: