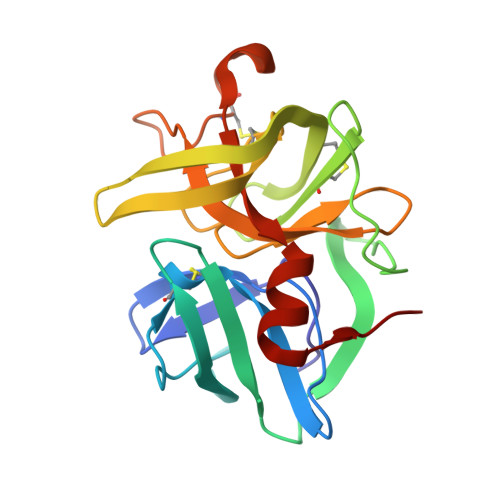
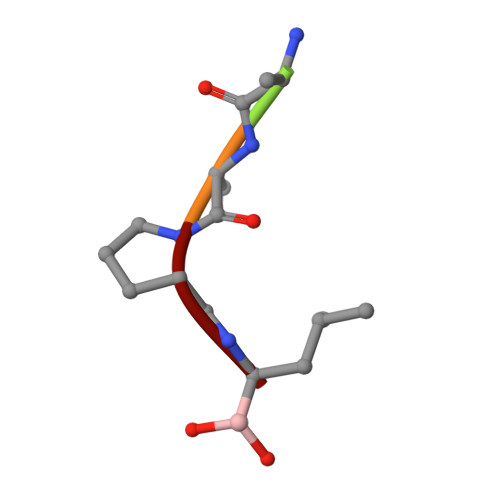
Structural basis for broad specificity in alpha-lytic protease mutants.
Bone, R., Fujishige, A., Kettner, C.A., Agard, D.A.(1991) Biochemistry 30: 10388-10398
- PubMed: 1931963 Search on PubMed
- DOI: https://doi.org/10.1021/bi00107a005
- Primary Citation Related Structures:
2LPR, 3LPR, 5LPR, 6LPR, 7LPR, 8LPR, 9LPR - PubMed Abstract:
Binding pocket mutants of alpha-lytic protease (Met 192----Ala and Met 213----Ala) have been constructed recently in an effort to create a protease specific for Met just prior to the scissile bond. Instead, mutation resulted in proteases with extraordinarily broad specificity profiles and high activity [Bone, R., Silen, J. L., & Agard, D. A. (1989) Nature 339, 191-195]. To understand the structural basis for the unexpected specificity profiles of these mutants, high-resolution X-ray crystal structures have been determined for complexes of each mutant with a series of systematically varying peptidylboronic acids. These inhibitory analogues of high-energy reaction intermediates provide models for how substrates with different side chains interact with the enzyme during the transition state. Fifteen structures have been analyzed qualitatively and quantitatively with respect to enzyme-inhibitor hydrogen-bond lengths, buried hydrophobic surface area, unfilled cavity volume, and the magnitude of inhibitor accommodating conformational adjustments (particularly in the region of another binding pocket residue, Val 217A). Comparison of these four parameters with the Ki of each inhibitor and the kcat and Km of the analogous substrates indicates that while no single structural parameter consistently correlates with activity or inhibition, the observed data can be understood as a combination of effects. Furthermore, the relative contribution of each term differs for the three enzymes, reflecting the altered conformational energetics of each mutant. From the extensive structural analysis, it is clear that enzyme flexibility, especially in the region of Val 217A, is primarily responsible for the exceptionally broad specificity observed in either mutant. Taken together, the observed patterns of substrate specificity can be understood to arise directly from interactions between the substrate and the residues lining the specificity pocket and indirectly from interactions between peripheral regions of the protein and the active-site region that serve to modulate active-site flexibility.
- Department of Biochemistry, Howard Hughes Medical Institute, University of California, San Francisco 94143-0448.
Organizational Affiliation: