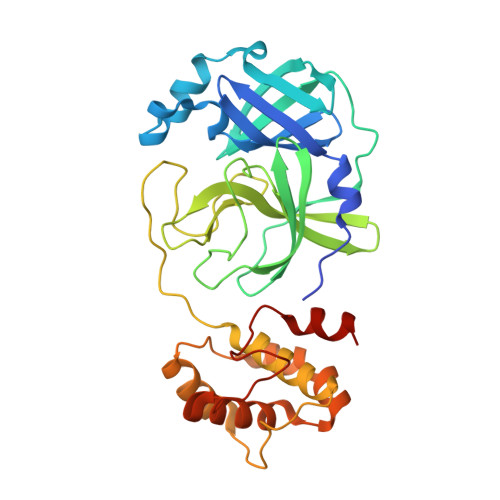
Structure-Guided Design of Conformationally Constrained Cyclohexane Inhibitors of Severe Acute Respiratory Syndrome Coronavirus-2 3CL Protease.
Dampalla, C.S., Kim, Y., Bickmeier, N., Rathnayake, A.D., Nguyen, H.N., Zheng, J., Kashipathy, M.M., Baird, M.A., Battaile, K.P., Lovell, S., Perlman, S., Chang, K.O., Groutas, W.C.(2021) J Med Chem 64: 10047-10058
- PubMed: 34213885 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.jmedchem.1c00319
- Primary Citation Related Structures:
7LKR, 7LKS, 7LKT, 7LKU, 7LKV, 7LKW, 7LKX - PubMed Abstract:
A series of nondeuterated and deuterated dipeptidyl aldehyde and masked aldehyde inhibitors that incorporate in their structure a conformationally constrained cyclohexane moiety was synthesized and found to potently inhibit severe acute respiratory syndrome coronavirus-2 3CL protease in biochemical and cell-based assays. Several of the inhibitors were also found to be nanomolar inhibitors of Middle East respiratory syndrome coronavirus 3CL protease. The corresponding latent aldehyde bisulfite adducts were found to be equipotent to the precursor aldehydes. High-resolution cocrystal structures confirmed the mechanism of action and illuminated the structural determinants involved in binding. The spatial disposition of the compounds disclosed herein provides an effective means of accessing new chemical space and optimizing pharmacological activity. The cellular permeability of the identified inhibitors and lack of cytotoxicity warrant their advancement as potential therapeutics for COVID-19.
- Department of Chemistry, Wichita State University, Wichita, Kansas 67260, United States.
Organizational Affiliation: