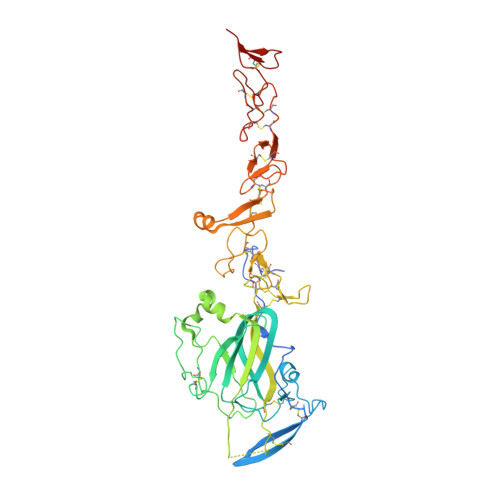
The dynamic nature of netrin-1 and the structural basis for glycosaminoglycan fragment-induced filament formation.
Meier, M., Gupta, M., Akgul, S., McDougall, M., Imhof, T., Nikodemus, D., Reuten, R., Moya-Torres, A., To, V., Ferens, F., Heide, F., Padilla-Meier, G.P., Kukura, P., Huang, W., Gerisch, B., Morgelin, M., Poole, K., Antebi, A., Koch, M., Stetefeld, J.(2023) Nat Commun 14: 1226-1226
- PubMed: 36869049 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-36692-w
- Primary Citation Related Structures:
7LER, 7LRF - PubMed Abstract:
Netrin-1 is a bifunctional chemotropic guidance cue that plays key roles in diverse cellular processes including axon pathfinding, cell migration, adhesion, differentiation, and survival. Here, we present a molecular understanding of netrin-1 mediated interactions with glycosaminoglycan chains of diverse heparan sulfate proteoglycans (HSPGs) and short heparin oligosaccharides. Whereas interactions with HSPGs act as platform to co-localise netrin-1 close to the cell surface, heparin oligosaccharides have a significant impact on the highly dynamic behaviour of netrin-1. Remarkably, the monomer-dimer equilibrium of netrin-1 in solution is abolished in the presence of heparin oligosaccharides and replaced with highly hierarchical and distinct super assemblies leading to unique, yet unknown netrin-1 filament formation. In our integrated approach we provide a molecular mechanism for the filament assembly which opens fresh paths towards a molecular understanding of netrin-1 functions.
- Department of Chemistry, University of Manitoba, Winnipeg, Canada.
Organizational Affiliation: