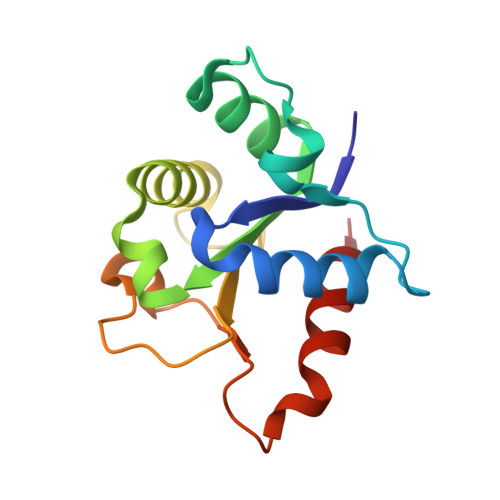
MyD88 TIR domain higher-order assembly interactions revealed by microcrystal electron diffraction and serial femtosecond crystallography.
Clabbers, M.T.B., Holmes, S., Muusse, T.W., Vajjhala, P.R., Thygesen, S.J., Malde, A.K., Hunter, D.J.B., Croll, T.I., Flueckiger, L., Nanson, J.D., Rahaman, M.H., Aquila, A., Hunter, M.S., Liang, M., Yoon, C.H., Zhao, J., Zatsepin, N.A., Abbey, B., Sierecki, E., Gambin, Y., Stacey, K.J., Darmanin, C., Kobe, B., Xu, H., Ve, T.(2021) Nat Commun 12: 2578-2578
- PubMed: 33972532 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-22590-6
- Primary Citation Related Structures:
7BEQ, 7BER, 7L6W - PubMed Abstract:
MyD88 and MAL are Toll-like receptor (TLR) adaptors that signal to induce pro-inflammatory cytokine production. We previously observed that the TIR domain of MAL (MAL TIR ) forms filaments in vitro and induces formation of crystalline higher-order assemblies of the MyD88 TIR domain (MyD88 TIR ). These crystals are too small for conventional X-ray crystallography, but are ideally suited to structure determination by microcrystal electron diffraction (MicroED) and serial femtosecond crystallography (SFX). Here, we present MicroED and SFX structures of the MyD88 TIR assembly, which reveal a two-stranded higher-order assembly arrangement of TIR domains analogous to that seen previously for MAL TIR . We demonstrate via mutagenesis that the MyD88 TIR assembly interfaces are critical for TLR4 signaling in vivo, and we show that MAL promotes unidirectional assembly of MyD88 TIR . Collectively, our studies provide structural and mechanistic insight into TLR signal transduction and allow a direct comparison of the MicroED and SFX techniques.
- Department of Materials and Environmental Chemistry, Stockholm University, Stockholm, Sweden.
Organizational Affiliation: