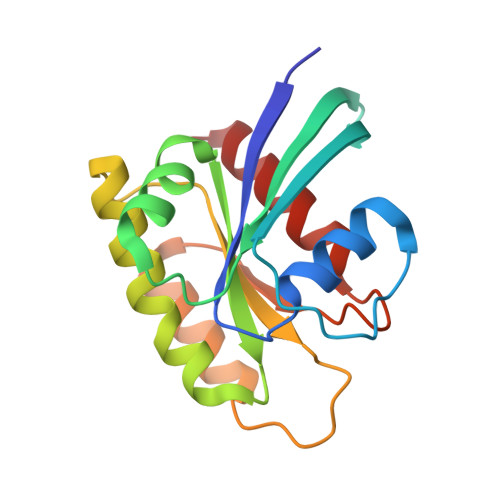
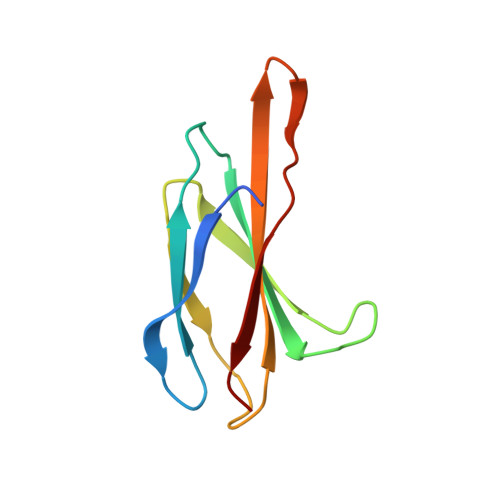
Selective and noncovalent targeting of RAS mutants for inhibition and degradation.
Teng, K.W., Tsai, S.T., Hattori, T., Fedele, C., Koide, A., Yang, C., Hou, X., Zhang, Y., Neel, B.G., O'Bryan, J.P., Koide, S.(2021) Nat Commun 12: 2656-2656
- PubMed: 33976200 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-22969-5
- Primary Citation Related Structures:
7L0F, 7L0G - PubMed Abstract:
Activating mutants of RAS are commonly found in human cancers, but to date selective targeting of RAS in the clinic has been limited to KRAS(G12C) through covalent inhibitors. Here, we report a monobody, termed 12VC1, that recognizes the active state of both KRAS(G12V) and KRAS(G12C) up to 400-times more tightly than wild-type KRAS. The crystal structures reveal that 12VC1 recognizes the mutations through a shallow pocket, and 12VC1 competes against RAS-effector interaction. When expressed intracellularly, 12VC1 potently inhibits ERK activation and the proliferation of RAS-driven cancer cell lines in vitro and in mouse xenograft models. 12VC1 fused to VHL selectively degrades the KRAS mutants and provides more extended suppression of mutant RAS activity than inhibition by 12VC1 alone. These results demonstrate the feasibility of selective targeting and degradation of KRAS mutants in the active state with noncovalent reagents and provide a starting point for designing noncovalent therapeutics against oncogenic RAS mutants.
- Perlmutter Cancer Center, New York University Langone Health, New York, NY, USA.
Organizational Affiliation: