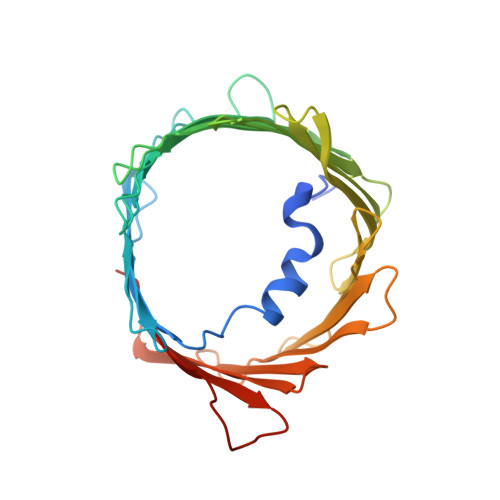
MicroED structure of lipid-embedded mammalian mitochondrial voltage-dependent anion channel.
Martynowycz, M.W., Khan, F., Hattne, J., Abramson, J., Gonen, T.(2020) Proc Natl Acad Sci U S A 117: 32380-32385
- PubMed: 33293416 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2020010117
- Primary Citation Related Structures:
7KUH - PubMed Abstract:
A structure of the murine voltage-dependent anion channel (VDAC) was determined by microcrystal electron diffraction (MicroED). Microcrystals of an essential mutant of VDAC grew in a viscous bicelle suspension, making it unsuitable for conventional X-ray crystallography. Thin, plate-like crystals were identified using scanning-electron microscopy (SEM). Crystals were milled into thin lamellae using a focused-ion beam (FIB). MicroED data were collected from three crystal lamellae and merged for completeness. The refined structure revealed unmodeled densities between protein monomers, indicative of lipids that likely mediate contacts between the proteins in the crystal. This body of work demonstrates the effectiveness of milling membrane protein microcrystals grown in viscous media using a focused ion beam for subsequent structure determination by MicroED. This approach is well suited for samples that are intractable by X-ray crystallography. To our knowledge, the presented structure is a previously undescribed mutant of the membrane protein VDAC, crystallized in a lipid bicelle matrix and solved by MicroED.
- Howard Hughes Medical Institute, University of California, Los Angeles, CA 90095.
Organizational Affiliation: