Antibodies That Engage the Hemagglutinin Receptor-Binding Site of Influenza B Viruses.
Bajic, G., Harrison, S.C.(2021) ACS Infect Dis 7: 1-5
- PubMed: 33274930 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsinfecdis.0c00726
- Primary Citation Related Structures:
7KQG, 7KQH, 7KQI - PubMed Abstract:
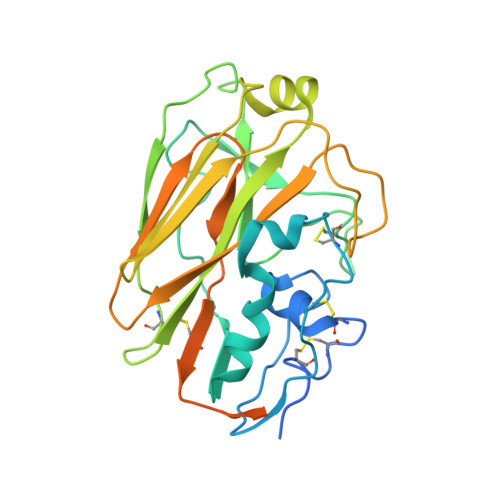
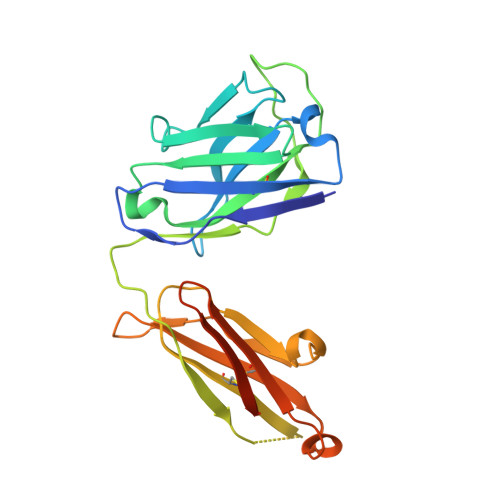
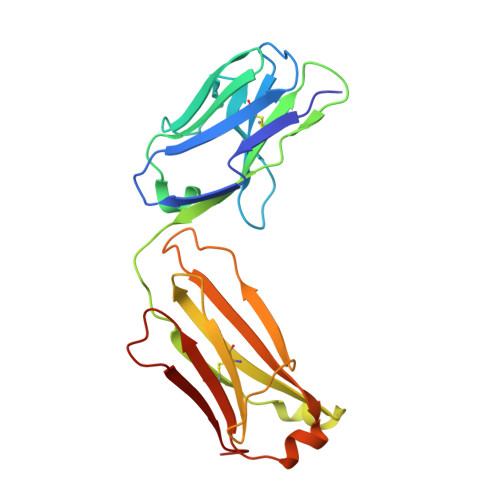
We describe cross-reactive human antibodies recognizing influenza B viruses spanning nearly 80 years of antigenic drift. Structures show that they engage the receptor-binding site (RBS) of the viral hemagglutinin with strong similarities to their influenza A counterparts, despite structural differences between the RBS of influenza A and B. Our data show that these antibodies readily cross-react with both influenza B Victoria and Yamagata lineages. We also note that all antibodies are encoded by IGHV3-9/IGK1-33. Future research will provide insight into the prevalence of these antibodies in the human population.
- Laboratory of Molecular Medicine, Boston Children's Hospital, Harvard Medical School, Boston, Massachusetts 02115, United States.
Organizational Affiliation: