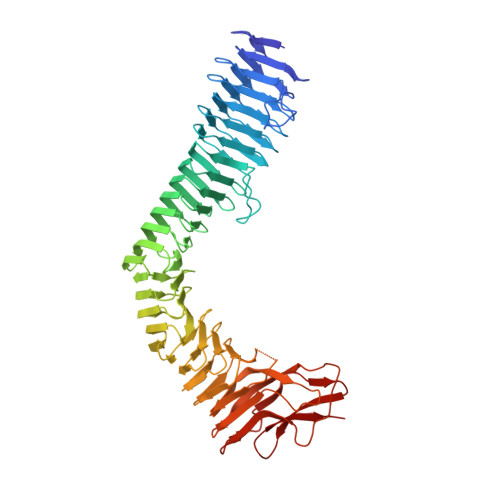
Variation of Antigen 43 self-association modulates bacterial compacting within aggregates and biofilms.
Vo, J.L., Ortiz, G.C.M., Totsika, M., Lo, A.W., Hancock, S.J., Whitten, A.E., Hor, L., Peters, K.M., Ageorges, V., Caccia, N., Desvaux, M., Schembri, M.A., Paxman, J.J., Heras, B.(2022) NPJ Biofilms Microbiomes 8: 20-20
- PubMed: 35396507 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41522-022-00284-1
- Primary Citation Related Structures:
7KO9, 7KOB, 7KOH - PubMed Abstract:
The formation of aggregates and biofilms enhances bacterial colonisation and infection progression by affording protection from antibiotics and host immune factors. Despite these advantages there is a trade-off, whereby bacterial dissemination is reduced. As such, biofilm development needs to be controlled to suit adaptation to different environments. Here we investigate members from one of largest groups of bacterial adhesins, the autotransporters, for their critical role in the assembly of bacterial aggregates and biofilms. We describe the structural and functional characterisation of autotransporter Ag43 variants from different Escherichia coli pathotypes. We show that specific interactions between amino acids on the contacting interfaces of adjacent Ag43 proteins drives a common mode of trans-association that leads to cell clumping. Furthermore, subtle variation of these interactions alters aggregation kinetics and the degree of compacting within cell clusters. Together, our structure-function investigation reveals an underlying molecular basis for variations in the density of bacterial communities.
- Department of Biochemistry and Chemistry, La Trobe Institute for Molecular Science, La Trobe University, Melbourne, VIC, 3086, Australia.
Organizational Affiliation: