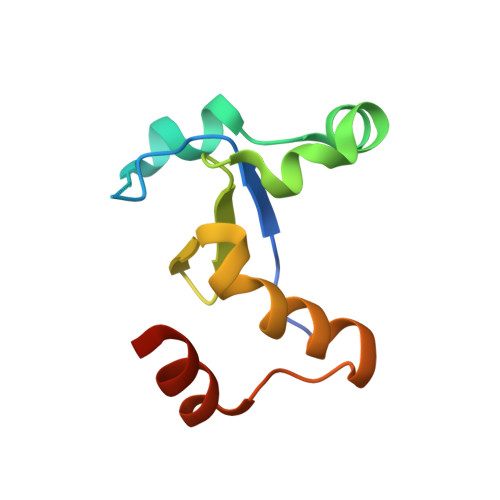
The middle lipin domain adopts a membrane-binding dimeric protein fold.
Gu, W., Gao, S., Wang, H., Fleming, K.D., Hoffmann, R.M., Yang, J.W., Patel, N.M., Choi, Y.M., Burke, J.E., Reue, K., Airola, M.V.(2021) Nat Commun 12: 4718-4718
- PubMed: 34354069 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-24929-5
- Primary Citation Related Structures:
7KIH, 7KIL, 7KIQ - PubMed Abstract:
Phospholipid synthesis and fat storage as triglycerides are regulated by lipin phosphatidic acid phosphatases (PAPs), whose enzymatic PAP function requires association with cellular membranes. Using hydrogen deuterium exchange mass spectrometry, we find mouse lipin 1 binds membranes through an N-terminal amphipathic helix, the Ig-like domain and HAD phosphatase catalytic core, and a middle lipin (M-Lip) domain that is conserved in mammalian and mammalian-like lipins. Crystal structures of the M-Lip domain reveal a previously unrecognized protein fold that dimerizes. The isolated M-Lip domain binds membranes both in vitro and in cells through conserved basic and hydrophobic residues. Deletion of the M-Lip domain in lipin 1 reduces PAP activity, membrane association, and oligomerization, alters subcellular localization, diminishes acceleration of adipocyte differentiation, but does not affect transcriptional co-activation. This establishes the M-Lip domain as a dimeric protein fold that binds membranes and is critical for full functionality of mammalian lipins.
- Department of Biochemistry and Cell Biology, Stony Brook University, Stony Brook, NY, USA.
Organizational Affiliation: