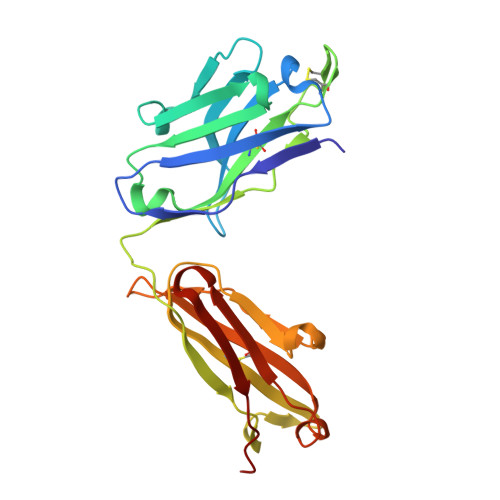
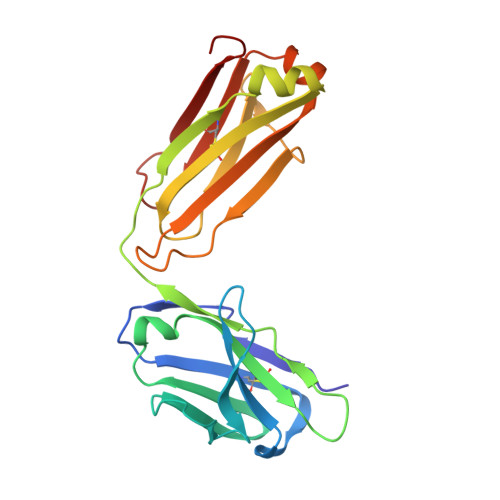
Ultrapotent human antibodies protect against SARS-CoV-2 challenge via multiple mechanisms.
Tortorici, M.A., Beltramello, M., Lempp, F.A., Pinto, D., Dang, H.V., Rosen, L.E., McCallum, M., Bowen, J., Minola, A., Jaconi, S., Zatta, F., De Marco, A., Guarino, B., Bianchi, S., Lauron, E.J., Tucker, H., Zhou, J., Peter, A., Havenar-Daughton, C., Wojcechowskyj, J.A., Case, J.B., Chen, R.E., Kaiser, H., Montiel-Ruiz, M., Meury, M., Czudnochowski, N., Spreafico, R., Dillen, J., Ng, C., Sprugasci, N., Culap, K., Benigni, F., Abdelnabi, R., Foo, S.C., Schmid, M.A., Cameroni, E., Riva, A., Gabrieli, A., Galli, M., Pizzuto, M.S., Neyts, J., Diamond, M.S., Virgin, H.W., Snell, G., Corti, D., Fink, K., Veesler, D.(2020) Science 370: 950-957
- PubMed: 32972994 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.abe3354
- Primary Citation Related Structures:
7K3Q, 7K43, 7K45, 7K4N - PubMed Abstract:
Efficient therapeutic options are needed to control the spread of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) that has caused more than 922,000 fatalities as of 13 September 2020. We report the isolation and characterization of two ultrapotent SARS-CoV-2 human neutralizing antibodies (S2E12 and S2M11) that protect hamsters against SARS-CoV-2 challenge. Cryo-electron microscopy structures show that S2E12 and S2M11 competitively block angiotensin-converting enzyme 2 (ACE2) attachment and that S2M11 also locks the spike in a closed conformation by recognition of a quaternary epitope spanning two adjacent receptor-binding domains. Antibody cocktails that include S2M11, S2E12, or the previously identified S309 antibody broadly neutralize a panel of circulating SARS-CoV-2 isolates and activate effector functions. Our results pave the way to implement antibody cocktails for prophylaxis or therapy, circumventing or limiting the emergence of viral escape mutants.
- Department of Biochemistry, University of Washington, Seattle, WA 98195, USA.
Organizational Affiliation: