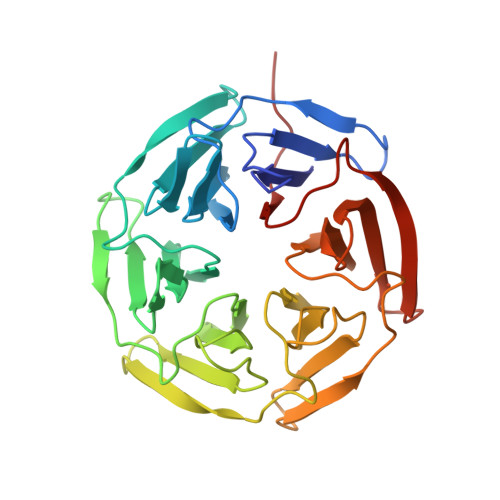
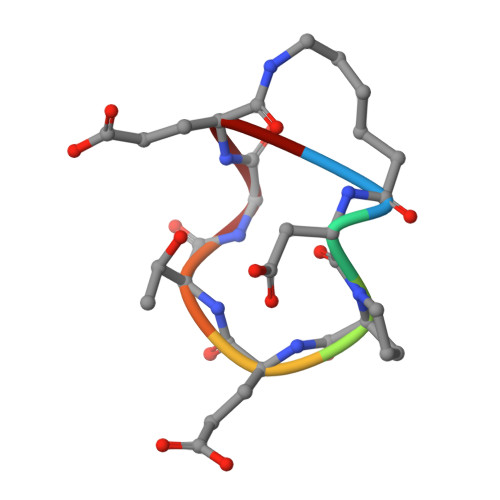
Recapitulating the Binding Affinity of Nrf2 for KEAP1 in a Cyclic Heptapeptide, Guided by NMR, X-ray Crystallography, and Machine Learning.
Ortet, P.C., Muellers, S.N., Viarengo-Baker, L.A., Streu, K., Szymczyna, B.R., Beeler, A.B., Allen, K.N., Whitty, A.(2021) J Am Chem Soc 143: 3779-3793
- PubMed: 33683866 Search on PubMed
- DOI: https://doi.org/10.1021/jacs.0c09799
- Primary Citation Related Structures:
7K28, 7K29, 7K2A, 7K2B, 7K2C, 7K2D, 7K2E, 7K2F, 7K2G, 7K2H, 7K2I, 7K2J, 7K2K, 7K2L, 7K2M, 7K2N, 7K2O, 7K2P, 7K2Q, 7K2R, 7K2S - PubMed Abstract:
Macrocycles, including macrocyclic peptides, have shown promise for targeting challenging protein-protein interactions (PPIs). One PPI of high interest is between Kelch-like ECH-Associated Protein-1 (KEAP1) and Nuclear Factor (Erythroid-derived 2)-like 2 (Nrf2). Guided by X-ray crystallography, NMR, modeling, and machine learning, we show that the full 20 nM binding affinity of Nrf2 for KEAP1 can be recapitulated in a cyclic 7-mer peptide, c[( D )-β-homoAla-DPETGE]. This compound was identified from the Nrf2-derived linear peptide GDEETGE ( K D = 4.3 μM) solely by optimizing the conformation of the cyclic compound, without changing any KEAP1 interacting residue. X-ray crystal structures were determined for each linear and cyclic peptide variant bound to KEAP1. Despite large variations in affinity, no obvious differences in the conformation of the peptide binding residues or in the interactions they made with KEAP1 were observed. However, analysis of the X-ray structures by machine learning showed that locations of strain in the bound ligand could be identified through patterns of subangstrom distortions from the geometry observed for unstrained linear peptides. We show that optimizing the cyclic peptide affinity was driven partly through conformational preorganization associated with a proline substitution at position 78 and with the geometry of the noninteracting residue Asp77 and partly by decreasing strain in the ETGE motif itself. This approach may have utility in dissecting the trade-off between conformational preorganization and strain in other ligand-receptor systems. We also identify a pair of conserved hydrophobic residues flanking the core DxETGE motif which play a conformational role in facilitating the high-affinity binding of Nrf2 to KEAP1.