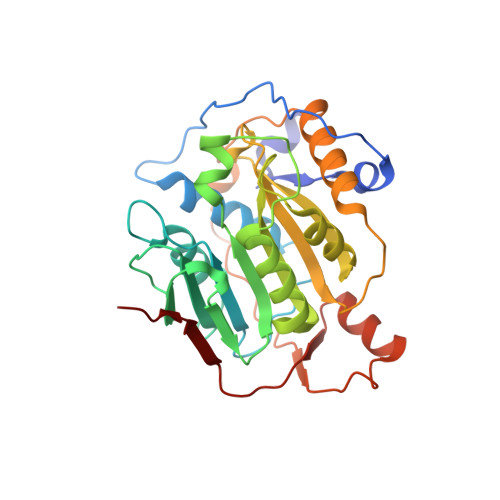
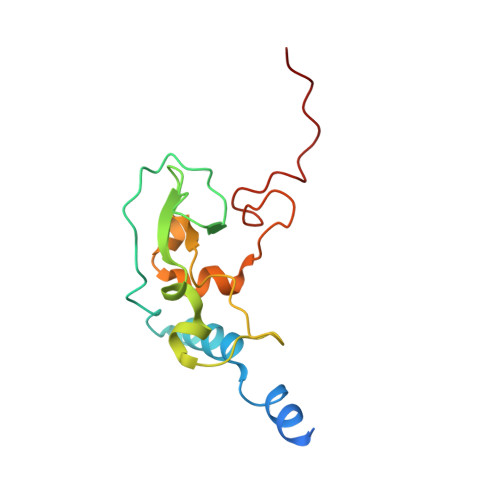
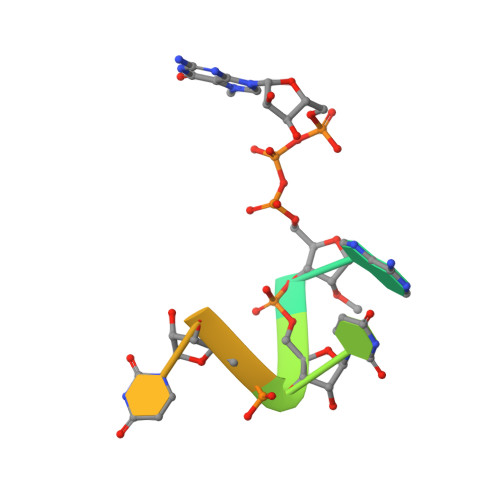
Mn2+ coordinates Cap-0-RNA to align substrates for efficient 2'-O-methyl transfer by SARS-CoV-2 nsp16
Minasov, G., Rosas-Lemus, M., Shuvalova, L., Inniss, N.L., Brunzelle, J.S., Daczkowski, C.M., Hoover, P., Mesecar, A.D., Satchell, K.J.F.(2021) bioRxiv