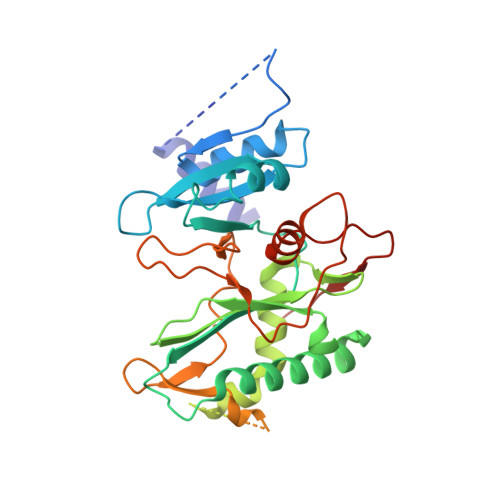
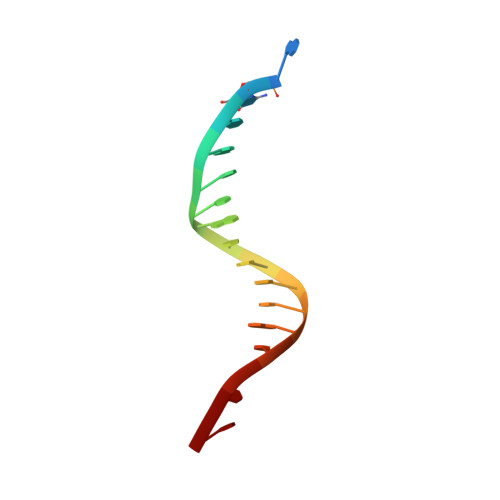
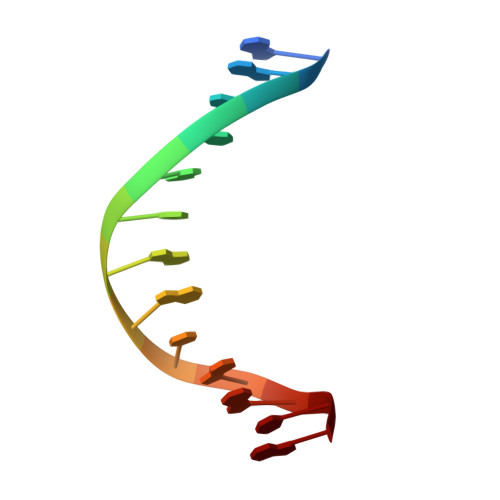
Structural basis of DNA synthesis opposite 8-oxoguanine by human PrimPol primase-polymerase.
Rechkoblit, O., Johnson, R.E., Gupta, Y.K., Prakash, L., Prakash, S., Aggarwal, A.K.(2021) Nat Commun 12: 4020-4020
- PubMed: 34188055 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-24317-z
- Primary Citation Related Structures:
7JK1, 7JKL, 7JKP, 7JL8, 7JLG - PubMed Abstract:
PrimPol is a human DNA polymerase-primase that localizes to mitochondria and nucleus and bypasses the major oxidative lesion 7,8-dihydro-8-oxoguanine (oxoG) via translesion synthesis, in mostly error-free manner. We present structures of PrimPol insertion complexes with a DNA template-primer and correct dCTP or erroneous dATP opposite the lesion, as well as extension complexes with C or A as a 3'-terminal primer base. We show that during the insertion of C and extension from it, the active site is unperturbed, reflecting the readiness of PrimPol to accommodate oxoG(anti). The misinsertion of A opposite oxoG(syn) also does not alter the active site, and is likely less favorable due to lower thermodynamic stability of the oxoG(syn)•A base-pair. During the extension step, oxoG(syn) induces an opening of its base-pair with A or misalignment of the 3'-A primer terminus. Together, the structures show how PrimPol accurately synthesizes DNA opposite oxidatively damaged DNA in human cells.
- Department of Pharmacological Sciences, Icahn School of Medicine at Mount Sinai, New York, NY, USA. olga.rechkoblit@mssm.edu.
Organizational Affiliation: