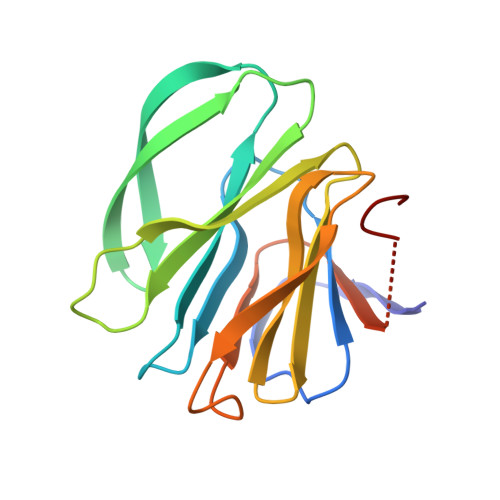
Structural basis of P[II] rotavirus evolution and host ranges under selection of histo-blood group antigens.
Xu, S., McGinnis, K.R., Liu, Y., Huang, P., Tan, M., Stuckert, M.R., Burnside, R.E., Jacob, E.G., Ni, S., Jiang, X., Kennedy, M.A.(2021) Proc Natl Acad Sci U S A 118
- PubMed: 34475219 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2107963118
- Primary Citation Related Structures:
6UT9, 6VKX, 7JHZ, 7KHU, 7KI5 - PubMed Abstract:
Group A rotaviruses cause severe gastroenteritis in infants and young children worldwide, with P[II] genogroup rotaviruses (RVs) responsible for >90% of global cases. RVs have diverse host ranges in different human and animal populations determined by host histo-blood group antigen (HBGA) receptor polymorphism, but details governing diversity, host ranges, and species barriers remain elusive. In this study, crystal structures of complexes of the major P[II] genogroup P[4] and P[8] genotype RV VP8* receptor-binding domains together with Lewis epitope-containing LNDFH I glycans in combination with VP8* receptor-glycan ligand affinity measurements based on NMR titration experiments revealed the structural basis for RV genotype-specific switching between ββ and βα HBGA receptor-binding sites that determine RV host ranges. The data support the hypothesis that P[II] RV evolution progressed from animals to humans under the selection of type 1 HBGAs guided by stepwise host synthesis of type 1 ABH and Lewis HBGAs. The results help explain disease burden, species barriers, epidemiology, and limited efficacy of current RV vaccines in developing countries. The structural data has the potential to impact the design of future vaccine strategies against RV gastroenteritis.
- Department of Chemistry and Biochemistry, Miami University, Oxford, OH 45056.
Organizational Affiliation: