Crystal Structure of a rat Autotaxin complex
Hunziker, D., Joachim, S.C., Ullmer, C., Rudolph, M.G.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Isoform 2 of Ectonucleotide pyrophosphatase/phosphodiesterase family member 2 | 846 | Rattus norvegicus | Mutation(s): 3 Gene Names: Enpp2, Atx, Npps2 EC: 3.1.4.39 (PDB Primary Data), 3.1.4.4 (UniProt) | 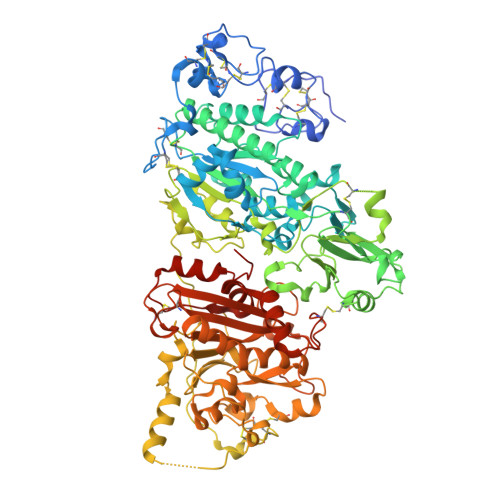 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q64610 | ||||
Glycosylation | |||||
| Glycosylation Sites: 1 | Go to GlyGen: Q64610-1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | D Interactions |
| alpha-D-mannopyranose-(1-2)-alpha-D-mannopyranose-(1-3)-alpha-D-mannopyranose-(1-6)-[alpha-D-mannopyranose-(1-2)-alpha-D-mannopyranose-(1-3)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | B | 8 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G33303XH GlyCosmos: G33303XH GlyGen: G33303XH | |||||
| Ligands 6 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| XHQ (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | D [auth A] | (3,5-dichlorophenyl)methyl (7R)-7-[(1H-benzotriazole-6-carbonyl)amino]-5-oxa-2-azaspiro[3.4]octane-2-carboxylate C21 H19 Cl2 N5 O4 OAHNIUIHQZJKPW-UHFFFAOYSA-N |  | ||
| MES Download:Ideal Coordinates CCD File | C [auth A] | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID C6 H13 N O4 S SXGZJKUKBWWHRA-UHFFFAOYSA-N |  | ||
| ZN Download:Ideal Coordinates CCD File | G [auth A] | ZINC ION Zn PTFCDOFLOPIGGS-UHFFFAOYSA-N |  | ||
| ACT Download:Ideal Coordinates CCD File | E [auth A] | ACETATE ION C2 H3 O2 QTBSBXVTEAMEQO-UHFFFAOYSA-M |  | ||
| CA Download:Ideal Coordinates CCD File | F [auth A], I [auth A] | CALCIUM ION Ca BHPQYMZQTOCNFJ-UHFFFAOYSA-N |  | ||
| NA Download:Ideal Coordinates CCD File | H [auth A], J [auth A] | SODIUM ION Na FKNQFGJONOIPTF-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 84.176 | α = 90 |
| b = 91.747 | β = 90 |
| c = 119.809 | γ = 90 |
| Software Name | Purpose |
|---|---|
| XSCALE | data scaling |
| REFMAC | refinement |
| PDB_EXTRACT | data extraction |
| XDS | data reduction |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| F. Hoffmann-La Roche LTD | Switzerland | -- |