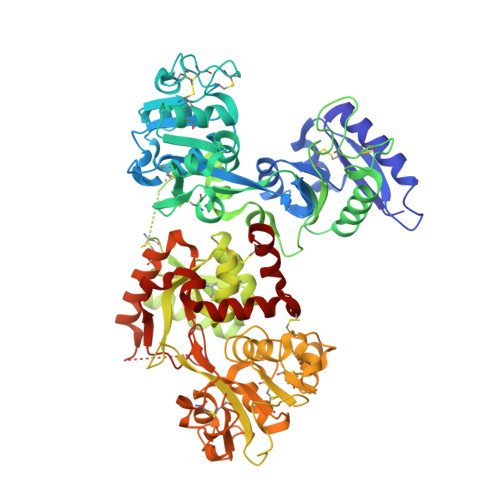
Binding of ruthenium and osmium at non‐iron sites of transferrin accounts for their iron-independent cellular uptake.
Wang, M., Wang, H., Xu, X., Lai, T.P., Zhou, Y., Hao, Q., Li, H., Sun, H.(2022) J Inorg Biochem 234: 111885-111885
- PubMed: 35690040 Search on PubMed
- DOI: https://doi.org/10.1016/j.jinorgbio.2022.111885
- Primary Citation Related Structures:
5WTD, 5X5P, 7FFM, 7FFU - PubMed Abstract:
Being identified with less toxic and generally showing selective effects for solid tumor metastases, ruthenium and osmium compounds are promising drug candidates for clinical uses. Human serum proteins, such as albumin and transferrin, play vital roles in the transportation and accumulation of ruthenium and osmium agents into target tissues. However, the molecular mechanism of how transferrin transport ruthenium and their osmium analogues at atomic level remains obscure. In this study, we uncovered that the cellular uptake of Os 3+ or Ru 3+ are not competed by Fe 3+ . To unveil the molecular mechanism behind the phenomena, we report the first crystal structures of human serum transferrin (hTF) in complex with ruthenium and osmium compounds bound to the non-conserved residues on the surface of hTF without altering its overall conformation. As for Ru 3+ and Os 3+ , these binding sites by descending affinity are: His14/His289, His349-350 ~ His578/Arg581. Ruthenium drugs and their osmium analogues preferentially bind to His14/His289 with bipyridine or imidazole ligands leaving. These binding sites on hTF surface are also available in human lactoferrin and some transferrin family member of other species. The presence of these binding sites makes the cellular uptake of Ru 3+ and Os 3+ less affected by Fe 3+ , compare to Zr 4+ or Hf 4+ . Collectively, these findings are critical for our understanding of the role of serum transferrin in cellular delivery of ruthenium and osmium anticancer agents.
- Department of Chemistry and CAS-HKU Joint Laboratory of Metallomics for Health and Environment, the University of Hong Kong, Pokfulam Road, Hong Kong SAR, China; School of Chemistry and Molecular Engineering, East China Normal University, No. 3663 Zhong Shan Road North, Shanghai 200062, China.
Organizational Affiliation: