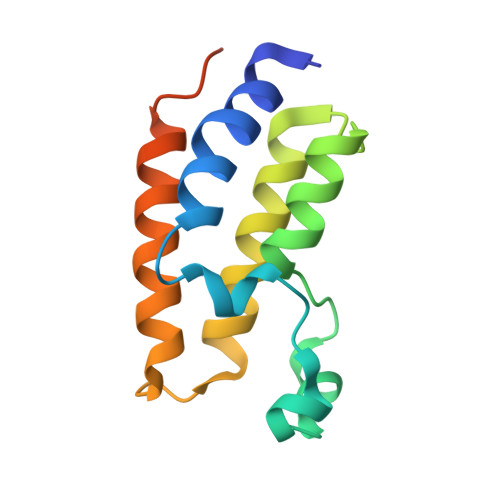
Structural investigation of a pyrano-1,3-oxazine derivative and the phenanthridinone core moiety against BRD2 bromodomains.
Arole, A.H., Deshmukh, P., Sridhar, A., Padmanabhan, B.(2022) Acta Crystallogr F Struct Biol Commun 78: 119-127
- PubMed: 35234137 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X22001066
- Primary Citation Related Structures:
7ENV, 7ENZ, 7EO5 - PubMed Abstract:
The BET (bromodomain and extra-terminal) family of proteins recognize the acetylated histone code on chromatin and play important roles in transcriptional co-regulation. BRD2 and BRD4, which belong to the BET family, are promising drug targets for the management of chronic diseases. The discovery of new scaffold molecules, a pyrano-1,3-oxazine derivative (NSC 328111; NS5) and phenanthridinone-based derivatives (L10 and its core moiety L10a), as inhibitors of BRD2 bromodomains BD1 and BD2, respectively, has recently been reported. The compound NS5 has a significant inhibitory effect on BRD2 in glioblastoma. Here, the crystal structure of BRD2 BD2 in complex with NS5, refined to 2.0 Å resolution, is reported. Moreover, as the previously reported crystal structures of the BD1-NS5 complex and the BD2-L10a complex possess moderate electron density corresponding to the respective ligands, the crystal structures of these complexes were re-evaluated using new X-ray data. Together with biochemical studies using wild-type BRD2 BD1 and BD2 and various mutants, it is confirmed that the pyrano-1,3-oxazine and phenanthridinone derivatives are indeed potent inhibitors of BRD2 bromodomains.
- Department of Biophysics, National Institute of Mental Health and Neuro Sciences (NIMHANS), Hosur Main Road, Bengaluru 560 029, India.
Organizational Affiliation: