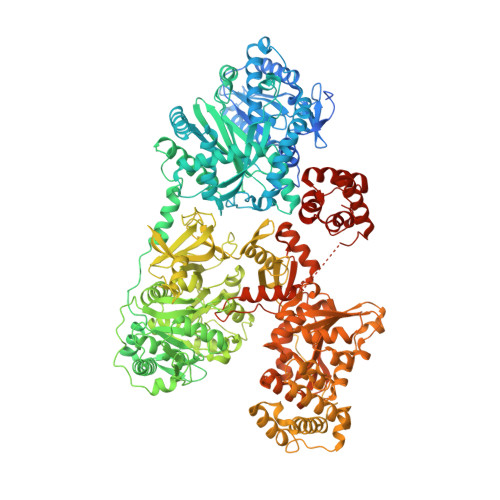
Catalytic trajectory of a dimeric nonribosomal peptide synthetase subunit with an inserted epimerase domain.
Wang, J., Li, D., Chen, L., Cao, W., Kong, L., Zhang, W., Croll, T., Deng, Z., Liang, J., Wang, Z.(2022) Nat Commun 13: 592-592
- PubMed: 35105906 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-28284-x
- Primary Citation Related Structures:
7EMY, 7EN1, 7EN2 - PubMed Abstract:
Nonribosomal peptide synthetases (NRPSs) are modular assembly-line megaenzymes that synthesize diverse metabolites with wide-ranging biological activities. The structural dynamics of synthetic elongation has remained unclear. Here, we present cryo-EM structures of PchE, an NRPS elongation module, in distinct conformations. The domain organization reveals a unique "H"-shaped head-to-tail dimeric architecture. The capture of both aryl and peptidyl carrier protein-tethered substrates and intermediates inside the heterocyclization domain and L-cysteinyl adenylate in the adenylation domain illustrates the catalytic and recognition residues. The multilevel structural transitions guided by the adenylation C-terminal subdomain in combination with the inserted epimerase and the conformational changes of the heterocyclization tunnel are controlled by two residues. Moreover, we visualized the direct structural dynamics of the full catalytic cycle from thiolation to epimerization. This study establishes the catalytic trajectory of PchE and sheds light on the rational re-engineering of domain-inserted dimeric NRPSs for the production of novel pharmaceutical agents.
- State Key Laboratory of Microbial Metabolism, Joint International Research Laboratory of Metabolic & Developmental Sciences, and School of Life Science & Biotechnology, Shanghai Jiao Tong University, Shanghai, China.
Organizational Affiliation: