Human p53 core domain with hot spot mutation R282W in complex with the natural CDKN1A(p21) p53-response element and Arsenic
Xing, Y.F., Lu, M.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Cellular tumor antigen p53 | 200 | Homo sapiens | Mutation(s): 1 Gene Names: TP53, P53 | 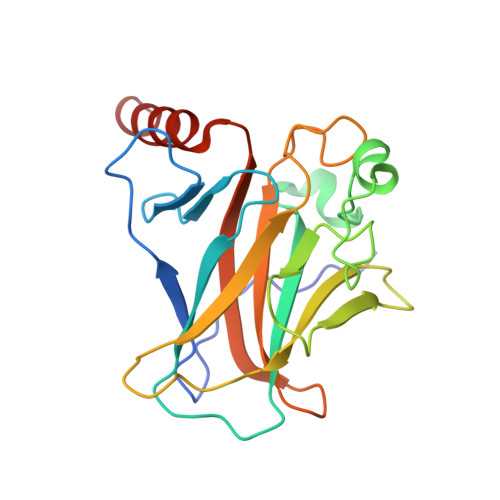 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P04637 GTEx: ENSG00000141510 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P04637 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| CDKN1A(p21) sense strand | 30 | Homo sapiens | 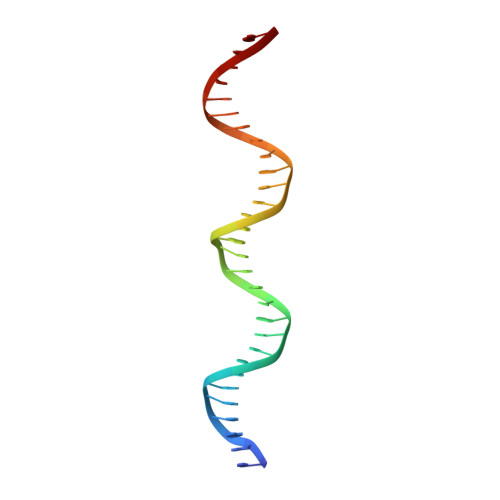 | |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
Entity ID: 3 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| CDKN1A(p21) anti-sense strand | 30 | Homo sapiens |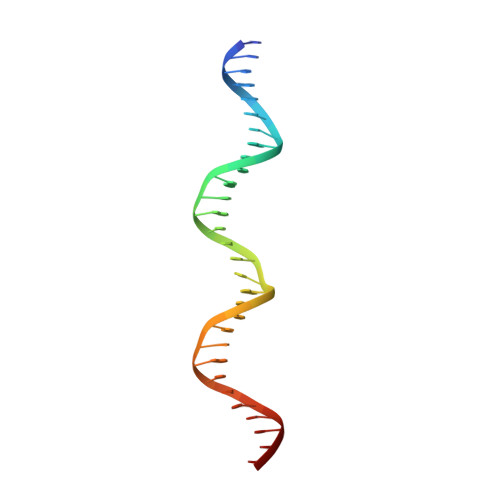  | |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| PEG Download:Ideal Coordinates CCD File | W [auth E], Z [auth F] | DI(HYDROXYETHYL)ETHER C4 H10 O3 MTHSVFCYNBDYFN-UHFFFAOYSA-N |  | ||
| ARS (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | AA [auth G] CA [auth H] M [auth A] O [auth B] Q [auth C] | ARSENIC As RBFQJDQYXXHULB-UHFFFAOYSA-N |  | ||
| ZN Download:Ideal Coordinates CCD File | BA [auth G] DA [auth H] N [auth A] P [auth B] R [auth C] | ZINC ION Zn PTFCDOFLOPIGGS-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 73.815 | α = 89.62 |
| b = 95.575 | β = 89.85 |
| c = 99.646 | γ = 75.79 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| Aimless | data scaling |
| PDB_EXTRACT | data extraction |
| XDS | data reduction |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Natural Science Foundation of China (NSFC) | China | 81861130368 |