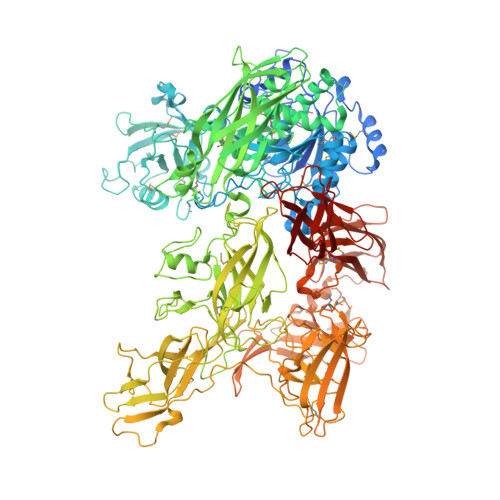
The Autocatalytic Cleavage Domain Is Not Required for the Activity of ScpC, a Virulence Protease from Streptococcus pyogenes : A Structural Insight.
Jobichen, C., Ying Chong, T., Hui Ling, T., Sivaraman, J.(2021) Biochemistry 60: 1564-1568
- PubMed: 33929828 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.1c00185
- Primary Citation Related Structures:
7EDD - PubMed Abstract:
Group A Streptococcus (GAS, or Streptococcus pyogenes ) is a leading human bacterial pathogen with diverse clinical manifestations, ranging from mild to life-threatening and to severe immune sequela. These diseases, combined, account for more than half a million deaths per year, globally. To accomplish its vast pathogenic potential, GAS expresses a multitude of virulent proteins, including the pivotal virulence factor ScpC. ScpC is a narrow-range surface-exposed subtilisin-like serine protease that cleaves the last 14 C-terminal amino acids of interleukin 8 (IL-8 or CXCL8) and impairs essential IL-8 signaling processes. As a result, neutrophil migration, bacterial killing, and the formation of neutrophil extracellular traps are strongly impaired. Also, ScpC has been identified as a potential vaccine candidate. ScpC undergoes an autocatalytic cleavage between Gln244 and Ser245, resulting in two polypeptide chains that assemble together forming the active protease. Previously, we reported that the region harboring the autocatalytic cleavage site, stretching from Gln213 to Asp272, is completely disordered. Here, we show that a deletion mutant (ScpC Δ60 ) of this region forms a single polypeptide chain, whose crystal structure we determined at 2.9 Å resolution. Moreover, we show that ScpC Δ60 is an active protease capable of cleaving its substrate IL-8 in a manner comparable to that of the wild type. These studies improve our understanding of the proteolytic activity of ScpC.
- Department of Biological Sciences, National University of Singapore, 14 Science Drive 4, Singapore 117543.
Organizational Affiliation: