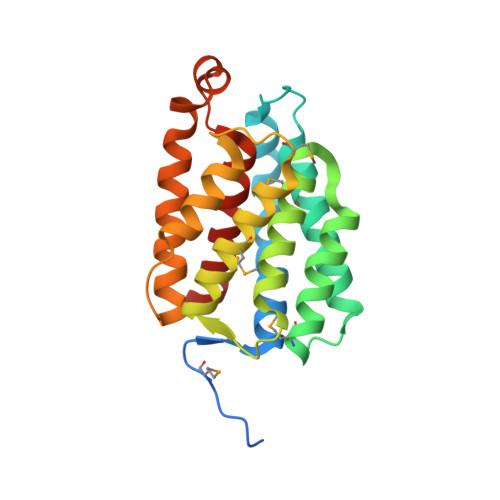
Crystal structure of a nucleotide-binding domain of fatty acid kinase FakA from Thermus thermophilus HB8.
Nakatani, M., Nakahara, S.Y., Fukui, K., Urano, M., Fujii, Y., Murakawa, T., Baba, S., Kumasaka, T., Okanishi, H., Kanai, Y., Yano, T., Masui, R.(2022) J Struct Biol 214: 107904-107904
- PubMed: 36228973 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2022.107904
- Primary Citation Related Structures:
7ED6, 7ED9 - PubMed Abstract:
Fatty acid kinase is necessary for the incorporation of exogenous fatty acids into membrane phospholipids. Fatty acid kinase consists of two components: a kinase component, FakA, that phosphorylates a fatty acid bound to a fatty acid-binding component, FakB. However, the molecular details underlying the phosphotransfer reaction remain to be resolved. We determined the crystal structure of the N-terminal domain of FakA bound to ADP from Thermus thermophilus HB8. The overall structure of this domain showed that the helical barrel fold is similar to the nucleotide-binding component of dihydroxyacetone kinase. The structure of the nucleotide-binding site revealed the roles of the conserved residues in recognition of ADP and Mg 2+ , but the N-terminal domain of FakA lacked the ADP-capping loop found in the dihydroxyacetone kinase component. Based on the structural similarity to the two subunits of dihydroxyacetone kinase complex, we constructed a model of the complex of T. thermophilus FakB and the N-terminal domain of FakA. In this model, the invariant Arg residue of FakB occupied a position that was spatially similar to that of the catalytically important Arg residue of dihydroxyacetone kinase, which predicted a composite active site in the Fatty acid kinase complex.
- Graduate School of Science, Osaka City University, 3-3-138 Sugimoto, Sumiyoshi-ku, Osaka 558-8585, Japan.
Organizational Affiliation: