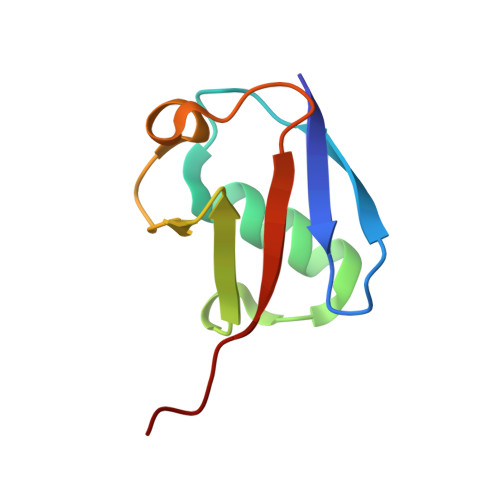
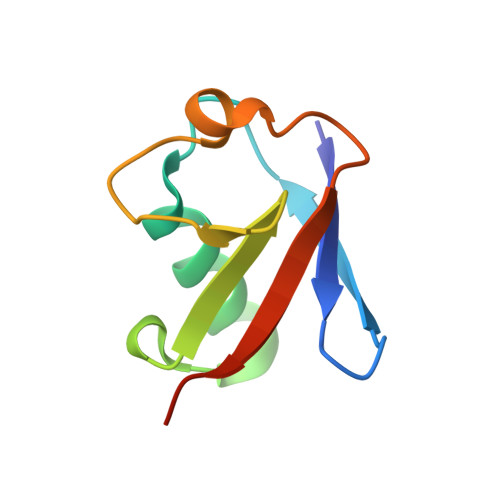
Structural basis for specific recognition of K6-linked polyubiquitin chains by the TAB2 NZF domain.
Li, Y., Okatsu, K., Fukai, S., Sato, Y.(2021) Biophys J 120: 3355-3362
- PubMed: 34242591 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bpj.2021.06.037
- Primary Citation Related Structures:
7E62 - PubMed Abstract:
TAK1-binding protein 2 (TAB2) has generally been considered to bind specifically to K63-linked polyubiquitin chains via its C-terminal Npl4 zinc-finger (NZF) domain. However, a recent study showed that the NZF domain of TAB2 (TAB2-NZF) could also interact with K6-linked polyubiquitin chains. Here, we report the crystal structure of TAB2-NZF in complex with K6-linked diubiquitin (K6-Ub 2 ) at 1.99-Å resolution. TAB2-NZF simultaneously interacts with the distal and proximal ubiquitin moieties of K6-Ub 2 . By comparing the structures of TAB2-NZF in complex with K6-Ub 2 and with K63-linked diubiquitin (K63-Ub 2 ), we reveal that the binding mechanism of TAB2-NZF with K6-Ub 2 is similar to that with K63-Ub 2 , except for the flexible C-terminal region of the distal ubiquitin. Therefore, we conclude that the C-terminal flexibility of the distal ubiquitin contributes to the dual specificity of TAB2-NZF toward K6- and K63-linked ubiquitin chains. This study provides important insights into the functions of K6-linked ubiquitin chains, which are currently unclear.
- Department of Computational Biology and Medical Sciences, Graduate School of Frontier Sciences, The University of Tokyo, Chiba, Japan.
Organizational Affiliation: