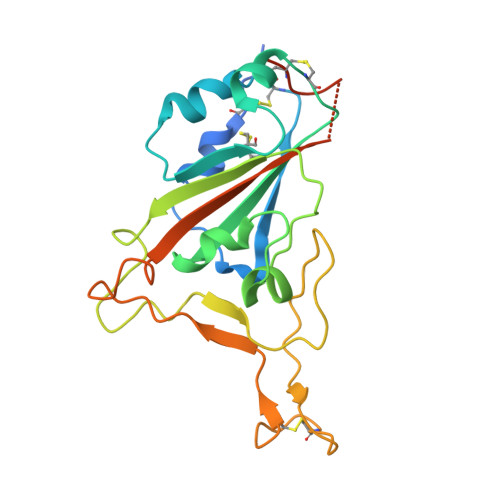
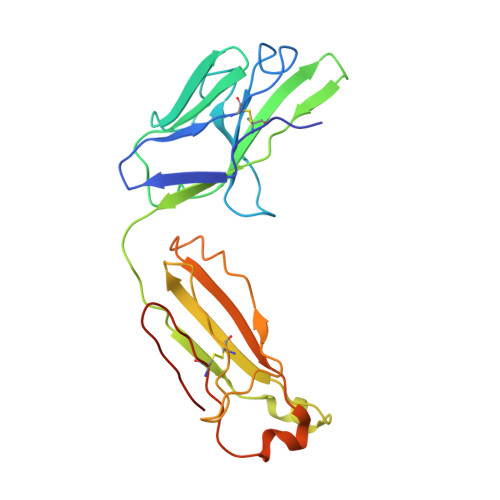
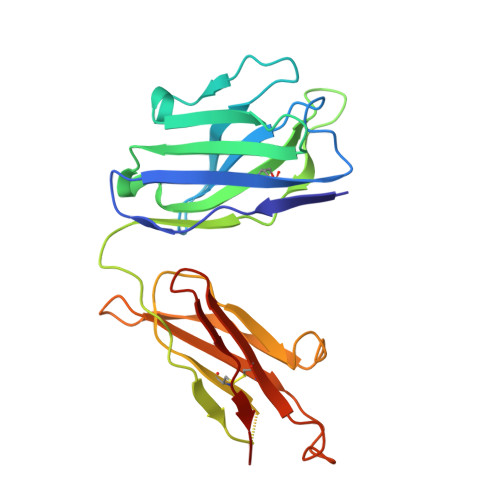
Screening of potent neutralizing antibodies against SARS-CoV-2 using convalescent patients-derived phage-display libraries.
Pan, Y., Du, J., Liu, J., Wu, H., Gui, F., Zhang, N., Deng, X., Song, G., Li, Y., Lu, J., Wu, X., Zhan, S., Jing, Z., Wang, J., Yang, Y., Liu, J., Chen, Y., Chen, Q., Zhang, H., Hu, H., Duan, K., Wang, M., Wang, Q., Yang, X.(2021) Cell Discov 7: 57-57
- PubMed: 34315862 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41421-021-00295-w
- Primary Citation Related Structures:
7E5Y - PubMed Abstract:
As the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) continues to threaten public health worldwide, the development of effective interventions is urgently needed. Neutralizing antibodies (nAbs) have great potential for the prevention and treatment of SARS-CoV-2 infection. In this study, ten nAbs were isolated from two phage-display immune libraries constructed from the pooled PBMCs of eight COVID-19 convalescent patients. Eight of them, consisting of heavy chains encoded by the immunoglobulin heavy-chain gene-variable region (IGHV)3-66 or IGHV3-53 genes, recognized the same epitope on the receptor-binding domain (RBD), while the remaining two bound to different epitopes. Among the ten antibodies, 2B11 exhibited the highest affinity and neutralization potency against the original wild-type (WT) SARS-CoV-2 virus (K D = 4.76 nM for the S1 protein, IC 50 = 6 ng/mL for pseudoviruses, and IC 50 = 1 ng/mL for authentic viruses), and potent neutralizing ability against B.1.1.7 pseudoviruses. Furthermore, 1E10, targeting a distinct epitope on RBD, exhibited different neutralization efficiency against WT SARS-CoV-2 and its variants B.1.1.7, B.1.351, and P.1. The crystal structure of the 2B11-RBD complexes revealed that the epitope of 2B11 highly overlaps with the ACE2-binding site. The in vivo experiment of 2B11 using AdV5-hACE2-transduced mice showed encouraging therapeutic and prophylactic efficacy against SARS-CoV-2. Taken together, our results suggest that the highly potent SARS-CoV-2-neutralizing antibody, 2B11, could be used against the WT SARS-CoV-2 and B.1.1.7 variant, or in combination with a different epitope-targeted neutralizing antibody, such as 1E10, against SARS-CoV-2 variants.
- National Engineering Technology Research Center for Combined Vaccines, Wuhan Institute of Biological Products Co. Ltd., Wuhan, Hubei, China.
Organizational Affiliation: