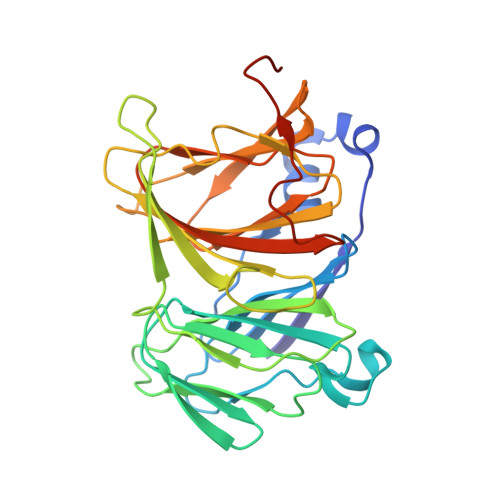
Crystal structures of Lacticaseibacillus 4-deoxy-L-threo-5-hexosulose-uronate ketol-isomerase KduI in complex with substrate analogs
Iwase, H., Yamamoto, Y., Yamada, A., Kawai, K., Oiki, S., Watanabe, D., Mikami, B., Takase, R., Hashimoto, W.(2023) J Appl Glycosci (1999)