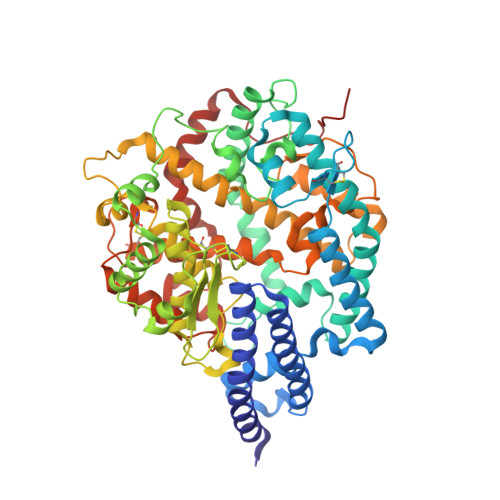
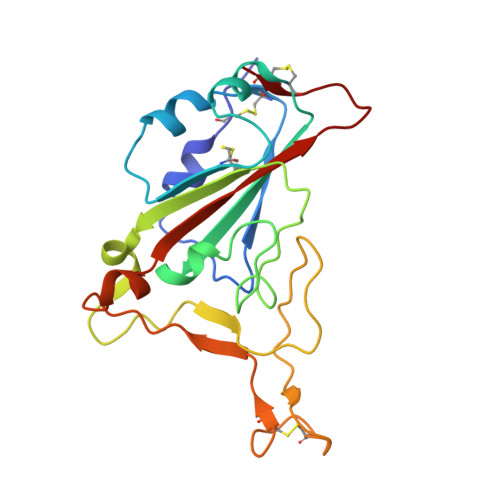
A proof of concept for neutralizing antibody-guided vaccine design against SARS-CoV-2.
Zhang, L., Cao, L., Gao, X.S., Zheng, B.Y., Deng, Y.Q., Li, J.X., Feng, R., Bian, Q., Guo, X.L., Wang, N., Qiu, H.Y., Wang, L., Cui, Z., Ye, Q., Chen, G., Lu, K.K., Chen, Y., Chen, Y.T., Pan, H.X., Yu, J., Yao, W., Zhu, B.L., Chen, J., Liu, Y., Qin, C.F., Wang, X., Zhu, F.C.(2021) Natl Sci Rev 8: nwab053-nwab053
- PubMed: 34676098 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nsr/nwab053
- Primary Citation Related Structures:
7D4G, 7DX4 - PubMed Abstract:
Mutations and transient conformational movements of the receptor binding domain (RBD) that make neutralizing epitopes momentarily unavailable present immune escape routes for severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). To mitigate viral escape, we developed a cocktail of neutralizing antibodies (NAbs) targeting epitopes located on different domains of spike (S) protein. Screening of a library of monoclonal antibodies generated from peripheral blood mononuclear cells of COVID-19 convalescent patients yielded potent NAbs, targeting the N-terminal domain (NTD) and RBD domain of S, effective at nM concentrations. Remarkably, a combination of RBD-targeting NAbs and NTD-binding NAbs, FC05, enhanced the neutralization potency in cell-based assays and an animal model. Results of competitive surface plasmon resonance assays and cryo-electron microscopy (cryo-EM) structures of antigen-binding fragments bound to S unveil determinants of immunogenicity. Combinations of immunogens, identified in the NTD and RBD of S, when immunized in rabbits and macaques, elicited potent protective immune responses against SARS-CoV-2. More importantly, two immunizations of this combination of NTD and RBD immunogens provided complete protection in macaques against a SARS-CoV-2 challenge, without observable antibody-dependent enhancement of infection. These results provide a proof of concept for neutralization-based immunogen design targeting SARS-CoV-2 NTD and RBD.
- National Health Commission of the People's Republic of China, Key Laboratory of Enteric Pathogenic Microbiology (Jiangsu Provincial Center for Disease Control and Prevention), Nanjing 210009, China.
Organizational Affiliation: