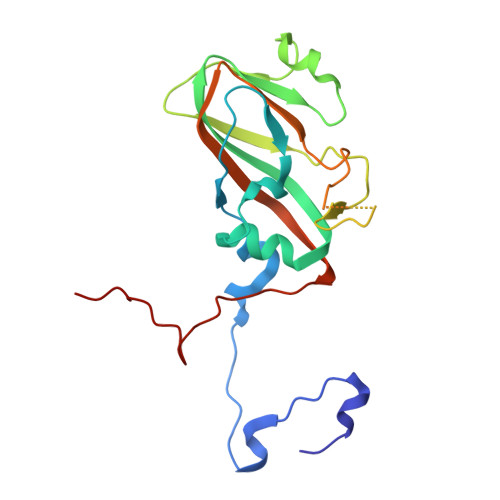
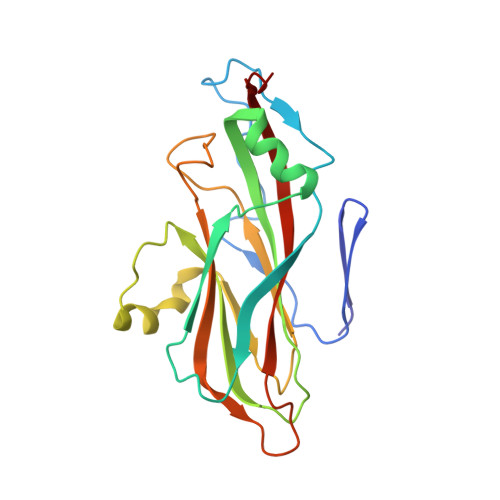
Structural and molecular basis for foot-and-mouth disease virus neutralization by two potent protective antibodies.
Dong, H., Liu, P., Bai, M., Wang, K., Feng, R., Zhu, D., Sun, Y., Mu, S., Li, H., Harmsen, M., Sun, S., Wang, X., Guo, H.(2022) Protein Cell 13: 446-453
- PubMed: 33599962 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s13238-021-00828-9
- Primary Citation Related Structures:
7DSS, 7DST - State Key Laboratory of Veterinary Etiological Biology and National Foot and Mouth Disease Reference Laboratory, Lanzhou Veterinary Research Institute, Chinese Academy of Agricultural Sciences, Lanzhou, 730046, China.
Organizational Affiliation: