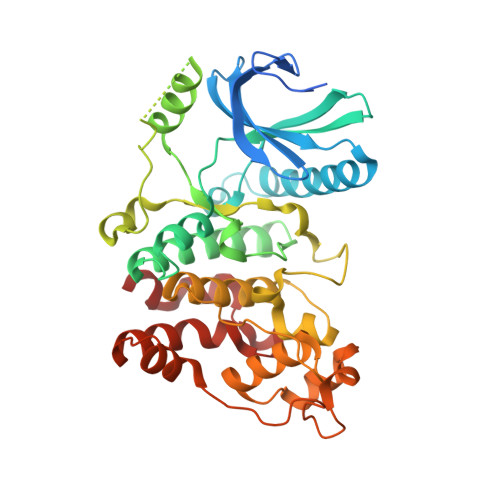
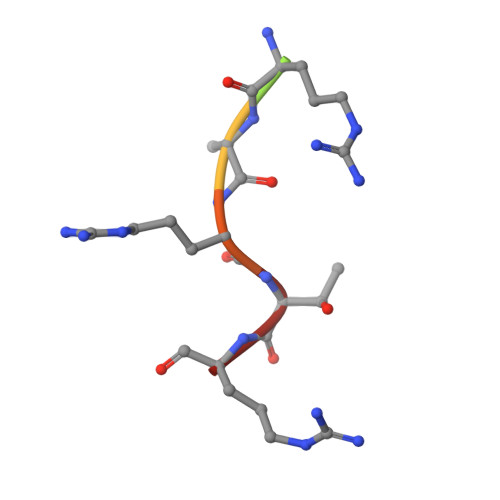
Protein-Protein Interaction Inhibitor of SRPKs Alters the Splicing Isoforms of VEGF and Inhibits Angiogenesis.
Li, Q., Zeng, C., Liu, H., Yung, K.W.Y., Chen, C., Xie, Q., Zhang, Y., Wan, S.W.C., Mak, B.S.W., Xia, J., Xiong, S., Ngo, J.C.K.(2021) iScience 24: 102423-102423
- PubMed: 33997701 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.isci.2021.102423
- Primary Citation Related Structures:
7DD1 - PubMed Abstract:
Serine-arginine (SR) protein kinases (SRPKs) regulate the functions of the SR-rich splicing factors by phosphorylating multiple serines within their C-terminal arginine-serine-rich domains. Dysregulation of these phosphorylation events has been implicated in many diseases, suggesting SRPKs are potential therapeutic targets. In particular, aberrant SRPK1 expression alters the balances of proangiogenic (VEGF 165 ) and antiangiogenic (VEGF 165 b) splicing isoforms of the key angiogenesis factor, vascular endothelial growth factor (VEGF), through the phosphorylation of prototypic SR protein SRSF1. Here, we report a protein-protein interaction (PPI) inhibitor of SRPKs, docking blocker of SRPK1 (DBS1), that specifically blocks a conserved substrate docking groove unique to SRPKs. DBS1 is a cell-permeable inhibitor that effectively inhibits the binding and phosphorylation of SRSF1 and subsequently switches VEGF splicing from the proangiogenic to the antiangiogenic isoform. Our findings thus provide a new direction for the development of SRPK inhibitors through targeting a unique PPI site to combat angiogenic diseases.
- School of Life Sciences, The Chinese University of Hong Kong, Shatin, N.T., Hong Kong SAR, China.
Organizational Affiliation: