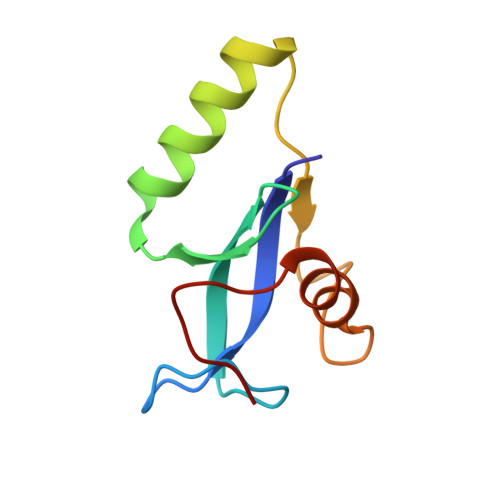
Structural and biochemical analyses of the tetrameric cell binding domain of Lys170 from enterococcal phage F170/08.
Xu, X., Zhang, D., Zhou, B., Zhen, X., Ouyang, S.(2021) Eur Biophys J 50: 721-729
- PubMed: 33609147 Search on PubMed
- DOI: https://doi.org/10.1007/s00249-021-01511-x
- Primary Citation Related Structures:
7D55 - PubMed Abstract:
Lysins are a class of hydrolytic enzymes used by bacteriophages to target and cleave the peptidoglycan of bacterial cell walls during their lytic cycle. The lysins from bacteriophages that infect Gram-positive bacteria are typically monomeric and consist of one or two catalytic domains (CD) and a cell binding domain (CBD). However, multimeric lysins encoded by a single gene have also been reported, among which Lys170 from enterococcal phage F170/08 was one of the first identified. Here, we determined the crystal structure of Lys170 CBD at 1.40 Å resolution. The structure reveals that Lys170 CBDs assemble into a tetrameric functional unit and that each monomer folds into a three-stranded β-sheet core capped on each side by an α-helix. In addition, we identified key residues of Lys170 CBD involved in host cell binding. Our work provides a basis for designing highly efficient lysins targeting Enterococcus faecalis.
- The Key Laboratory of Innate Immune Biology of Fujian Province, Provincial University Key Laboratory of Cellular Stress Response and Metabolic Regulation, Biomedical Research Center of South China, Key Laboratory of OptoElectronic Science and Technology for Medicine of the Ministry of Education, College of Life Sciences, Fujian Normal University, Fuzhou, China.
Organizational Affiliation: