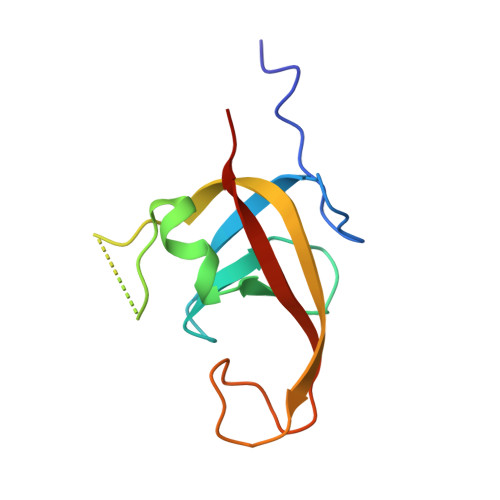
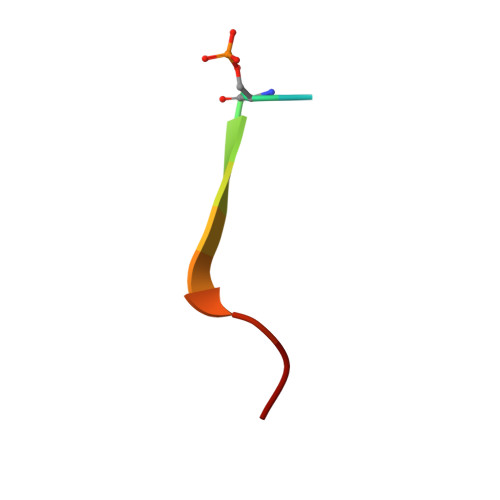
Phosphorylation regulates the binding of autophagy receptors to FIP200 Claw domain for selective autophagy initiation.
Zhou, Z., Liu, J., Fu, T., Wu, P., Peng, C., Gong, X., Wang, Y., Zhang, M., Li, Y., Wang, Y., Xu, X., Li, M., Pan, L.(2021) Nat Commun 12: 1570-1570
- PubMed: 33692357 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-21874-1
- Primary Citation Related Structures:
7CZG, 7CZM, 7D0E - PubMed Abstract:
The ULK complex initiates the autophagosome formation, and has recently been implicated in selective autophagy by interacting with autophagy receptors through its FIP200 subunit. However, the structural mechanism underlying the interactions of autophagy receptors with FIP200 and the relevant regulatory mechanism remain elusive. Here, we discover that the interactions of FIP200 Claw domain with autophagy receptors CCPG1 and Optineurin can be regulated by the phosphorylation in their respective FIP200-binding regions. We determine the crystal structures of FIP200 Claw in complex with the phosphorylated CCPG1 and Optineurin, and elucidate the detailed molecular mechanism governing the interactions of FIP200 Claw with CCPG1 and Optineurin as well as their potential regulations by kinase-mediated phosphorylation. In addition, we define the consensus FIP200 Claw-binding motif, and find other autophagy receptors that contain this motif within their conventional LC3-interacting regions. In all, our findings uncover a general and phosphoregulatable binding mode shared by many autophagy receptors to interact with FIP200 Claw for autophagosome biogenesis, and are valuable for further understanding the molecular mechanism of selective autophagy.
- State Key Laboratory of Bioorganic and Natural Products Chemistry, Center for Excellence in Molecular Synthesis, Shanghai Institute of Organic Chemistry, University of Chinese Academy of Sciences, Chinese Academy of Sciences, Shanghai, China.
Organizational Affiliation: