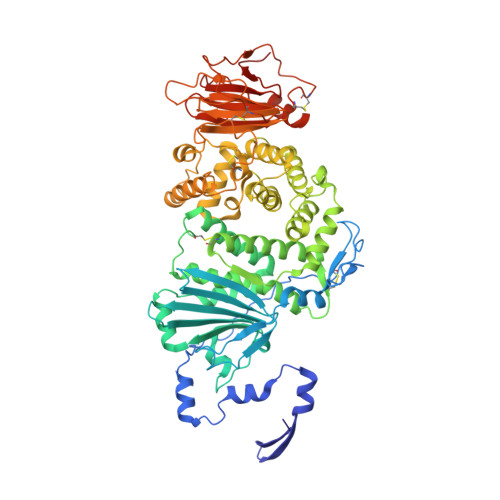
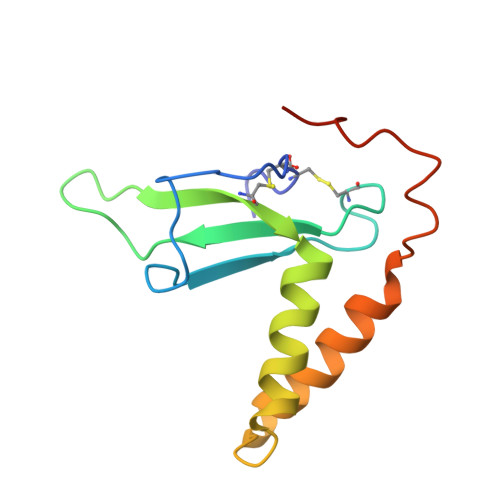
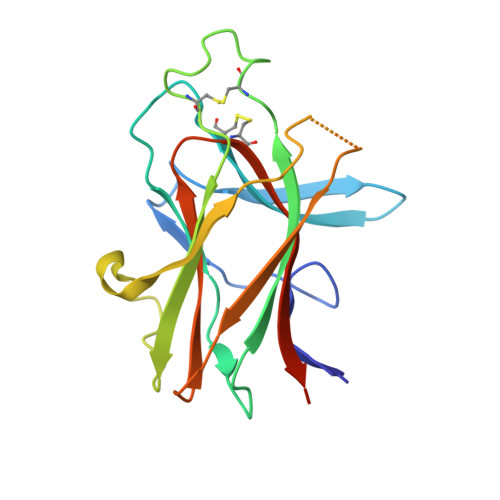
Molecular basis of EphA2 recognition by gHgL from gammaherpesviruses.
Su, C., Wu, L., Chai, Y., Qi, J., Tan, S., Gao, G.F., Song, H., Yan, J.(2020) Nat Commun 11: 5964-5964
- PubMed: 33235207 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-19617-9
- Primary Citation Related Structures:
7CZE, 7CZF - PubMed Abstract:
The human γ-herpesviruses Kaposi sarcoma associated herpesvirus (KSHV) and Epstein-Barr virus (EBV) are associated with many human malignancies. Viral glycoprotein H (gH) and glycoprotein L (gL) are crucial for the cell tropism by binding to specific receptors. Recently, EphA2 was identified as the specific entry receptor for both KSHV and EBV. Here, we characterized the crystal structures of KSHV gHgL or EBV gHgL in complex with the ligand binding domain (LBD) of EphA2. Both KSHV and EBV gHgL bind to the channel and peripheral regions of LBD primarily using gL. Extensive interactions with more contacts contribute to the higher affinity of KSHV gHgL to LBD than that of EBV gHgL. These binding characteristics were verified using cell-based fusion assays with mutations in key EphA2 residues. Our experiments suggest that multiple animal γ-herpesviruses could use EphA2 as an entry receptor, implying a potential threat to human health.
- College of Veterinary Medicine, China Agricultural University, Beijing, 100193, China.
Organizational Affiliation: