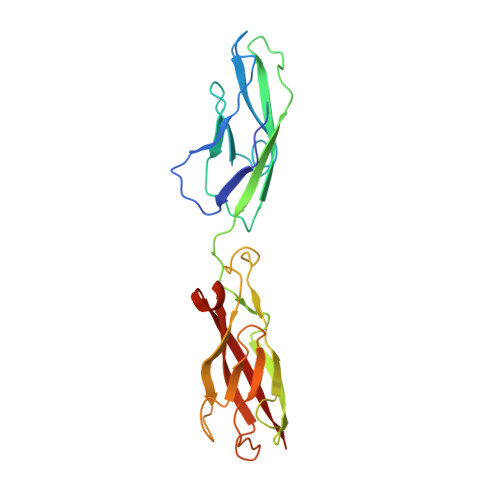
Regulation of cadherin dimerization by chemical fragments as a trigger to inhibit cell adhesion
Senoo, A., Ito, S., Nagatoishi, S., Saito, Y., Ueno, G., Kuroda, D., Yoshida, K., Tashima, T., Kudo, S., Sando, S., Tsumoto, K.(2021) Commun Biol 4: 1041