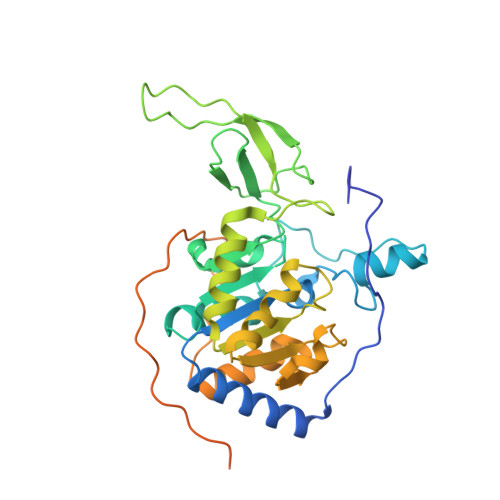
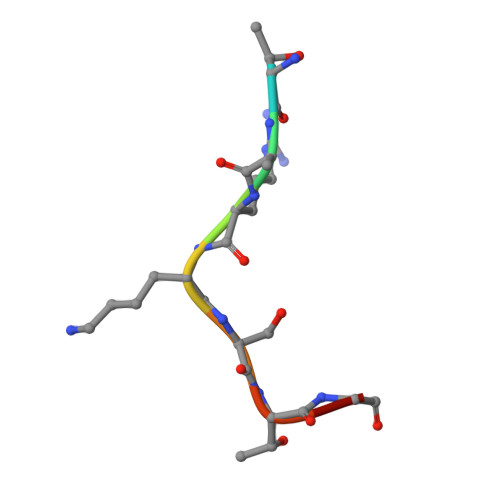
Reply to: Binding site for MDL-801 on SIRT6.
Huang, Z., Zhao, J., Deng, W., Chen, Y., Shang, J., Song, K., Zhang, L., Wang, C., Lu, S., Yang, X., He, B., Min, J., Hu, H., Tan, M., Xu, J., Zhang, Q., Zhong, J., Sun, X., Mao, Z., Lin, H., Xiao, M., Chin, Y.E., Jiang, H., Shen, H., Xu, Y., Chen, G., Zhang, J.(2021) Nat Chem Biol 17: 522-523
- PubMed: 33649600 Search on PubMed
- DOI: https://doi.org/10.1038/s41589-021-00750-5
- Primary Citation Related Structures:
7CL0, 7CL1 - Department of Pathophysiology, Shanghai Jiaotong University School of Medicine, Shanghai, China.
Organizational Affiliation: