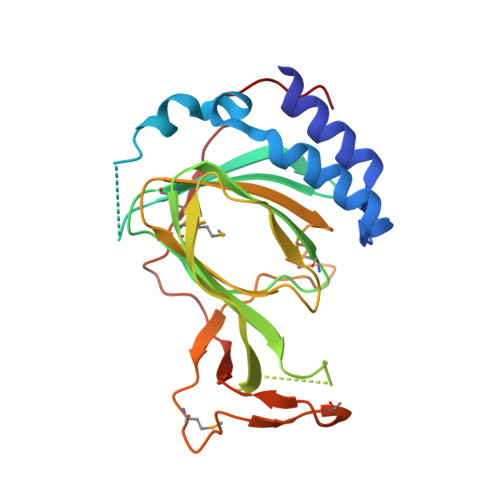
Molecular basis for cysteine oxidation by plant cysteine oxidases from Arabidopsis thaliana.
Chen, Z., Guo, Q., Wu, G., Wen, J., Liao, S., Xu, C.(2021) J Struct Biol 213: 107663-107663
- PubMed: 33207269 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2020.107663
- Primary Citation Related Structures:
7CHI, 7CHJ, 7CXZ - PubMed Abstract:
Plant Cysteine Oxidases (PCOs) play important roles in controlling the stability of Group VII ethylene response factors (ERF-VIIs) via Arg/N-degron pathway through catalyzing the oxidation of their N-Cys for subsequent Arginyl-tRNA--protein transferase 1 (ATE1) mediated arginine installation. Here we presented the crystal structures of PCO2, PCO4, and PCO5 from Arabidopsis thaliana (AtPCOs) and examined their in vitro activity by Mass spectrometry (MS). On the basis of Tris-bound AtPCO2, we modelled the structure of Cys-bound AtPCO2 and identified key AtPCO2 residues involved in N-Cys recognition and oxidation. Alanine substitution of potential N-Cys interaction residues impaired the activity of AtPCO5 remarkably. The structural research, complemented by mutagenesis and MS experiments, not only uncovers the substrate recognition and catalytic mode by AtPCOs, but also sheds light on the future design of potent inhibitors for plant cysteine oxidases.
- The First Affiliated Hospital of University of Science and Technology of China (Anhui Provincial Hospital), 230000 Hefei, China; MOE Key Laboratory for Membraneless Organelles and Cellular Dynamics, Hefei National Laboratory for Physical Sciences at the Microscale, University of Science and Technology of China, Hefei 230027, China.
Organizational Affiliation: