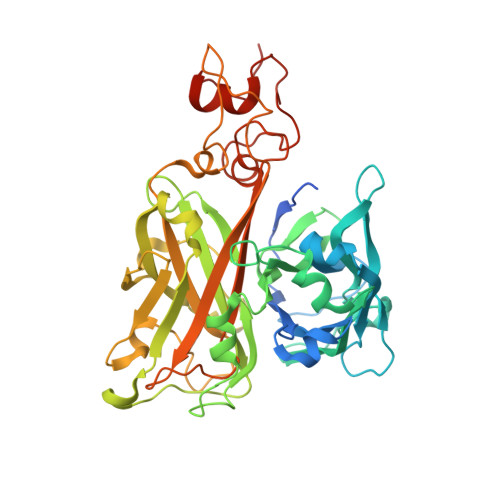
Structural mechanism for modulation of functional amyloid and biofilm formation by Staphylococcal Bap protein switch.
Ma, J., Cheng, X., Xu, Z., Zhang, Y., Valle, J., Fan, S., Zuo, X., Lasa, I., Fang, X.(2021) EMBO J 40: e107500-e107500
- PubMed: 34046916 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embj.2020107500
- Primary Citation Related Structures:
7C7R, 7C7U - PubMed Abstract:
The Staphylococcal Bap proteins sense environmental signals (such as pH, [Ca 2+ ]) to build amyloid scaffold biofilm matrices via unknown mechanisms. We here report the crystal structure of the aggregation-prone region of Staphylococcus aureus Bap which adopts a dumbbell-shaped fold. The middle module (MM) connecting the N-terminal and C-terminal lobes consists of a tandem of novel double-Ca 2+ -binding motifs involved in cooperative interaction networks, which undergoes Ca 2+ -dependent order-disorder conformational switches. The N-terminal lobe is sufficient to mediate amyloid aggregation through liquid-liquid phase separation and maturation, and subsequent biofilm formation under acidic conditions. Such processes are promoted by disordered MM at low [Ca 2+ ] but inhibited by ordered MM stabilized by Ca 2+ binding, with inhibition efficiency depending on structural integrity of the interaction networks. These studies illustrate a novel protein switch in pathogenic bacteria and provide insights into the mechanistic understanding of Bap proteins in modulation of functional amyloid and biofilm formation, which could be implemented in the anti-biofilm drug design.
- Beijing Advanced Innovation Center for Structural Biology, School of Life Sciences, Tsinghua University, Beijing, China.
Organizational Affiliation: