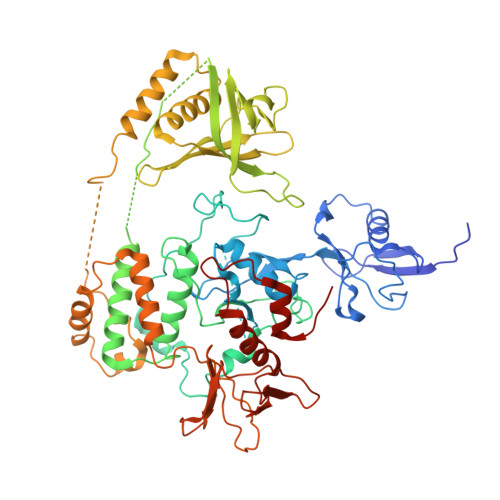
Structural basis of human full-length kindlin-3 homotrimer in an auto-inhibited state.
Bu, W., Levitskaya, Z., Loh, Z.Y., Jin, S., Basu, S., Ero, R., Yan, X., Wang, M., Ngan, S.F.C., Sze, S.K., Tan, S.M., Gao, Y.G.(2020) PLoS Biol 18: e3000755-e3000755
- PubMed: 32644996 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pbio.3000755
- Primary Citation Related Structures:
7C3M - PubMed Abstract:
Kindlin-1, -2, and -3 directly bind integrin β cytoplasmic tails to regulate integrin activation and signaling. Despite their functional significance and links to several diseases, structural information on full-length kindlin proteins remains unknown. Here, we report the crystal structure of human full-length kindlin-3, which reveals a novel homotrimer state. Unlike kindlin-3 monomer, which is the major population in insect and mammalian cell expression systems, kindlin-3 trimer does not bind integrin β cytoplasmic tail as the integrin-binding pocket in the F3 subdomain of 1 protomer is occluded by the pleckstrin homology (PH) domain of another protomer, suggesting that kindlin-3 is auto-inhibited upon trimer formation. This is also supported by functional assays in which kindlin-3 knockout K562 erythroleukemia cells reconstituted with the mutant kindlin-3 containing trimer-disrupting mutations exhibited an increase in integrin-mediated adhesion and spreading on fibronectin compared with those reconstituted with wild-type kindlin-3. Taken together, our findings reveal a novel mechanism of kindlin auto-inhibition that involves its homotrimer formation.
- School of Biological Sciences, Nanyang Technological University, Singapore, Singapore.
Organizational Affiliation: