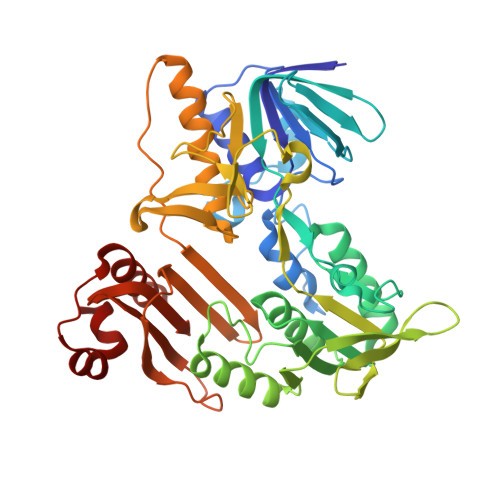
Biochemical and structural characterization of a robust and thermostable ascorbate recycling monodehydroascorbate reductase (MDHAR) from stress adapted pearl millet.
Sonkar, K.S., Achary, V.M.M., Sahoo, S., Reddy, M.K., Arockiasamy, A.(2023) Biochem Biophys Res Commun 662: 135-141
- PubMed: 37119729 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2023.04.034
- Primary Citation Related Structures:
7BQ6, 7BVI - PubMed Abstract:
Ascorbate (AsA) is a crucial antioxidant in plants, and its recycling is necessary for protecting cells from oxidative damage and imparting stress tolerance. The monodehydroascorbate reductase (MDHAR) enzyme of the ascorbate-glutathione pathway plays a vital role in recycling AsA from monodehydroascorbate (MDHA) radical. Pennisetum glaucum (Pg), also known as pearl millet, is known to be more tolerant to abiotic stress than other food crops, such as rice. However, the contribution of MDHAR from this sessile plant to its unique stress tolerance mechanism is not well understood. In this study, we isolated a gene encoding the MDHAR enzyme from heat stress-adapted pearl millet and characterized it using enzyme kinetics, thermal stability assays, and crystal structure determination. Our results indicate that PgMDHAR is a more robust enzyme than its rice counterpart (Oryza sativa; Os). We solved the crystal structure of PgMDHAR at 1.8 Å and found that the enzyme has a more compact structure and greater stability than OsMDHAR. Using hybrid quantum mechanics and molecular mechanics calculations, we demonstrate that the structure of PgMDHAR contributes to increased stability towards bound FAD. Overall, the higher structural stability and affinity for NADH demonstrated by PgMDHAR are expected to impart improved stress tolerance. Our findings suggest that transgenic food crops expressing MDHAR from stress-adapted pearl millet may exhibit better tolerance to oxidative stress in the unpredictable climatic conditions prevalent today.
- Membrane Protein Biology Group, ICGEB, Aruna Asaf Ali Marg, New Delhi, 110067, India.
Organizational Affiliation: