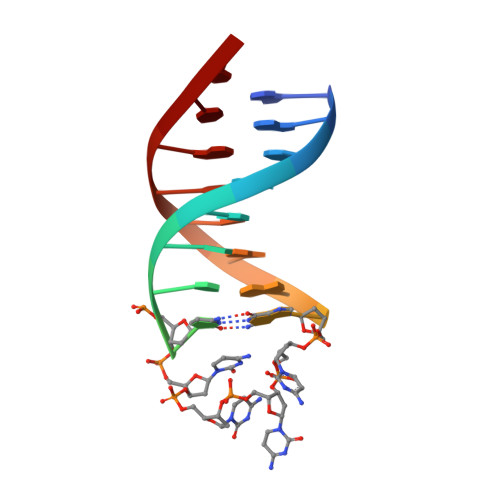
Stable Hairpin Structures Formed by Xylose-Based Nucleic Acids.
Mattelaer, C.A., Maiti, M., Smets, L., Maiti, M., Schepers, G., Mattelaer, H.P., Rosemeyer, H., Herdewijn, P., Lescrinier, E.(2021) Chembiochem 22: 1638-1645
- PubMed: 33427360 Search on PubMed
- DOI: https://doi.org/10.1002/cbic.202000803
- Primary Citation Related Structures:
7BFS, 7BFX - PubMed Abstract:
Xenobiology explores synthetic nucleic acid polymers as alternative carriers of genetic information to expand the central dogma. The xylo- and deoxyxylo-nucleic acids (XyNA and dXyNA), containing 3' epimers of riboses and deoxyriboses, are considered to be potential candidates for an orthogonal system. In this study, thermal and spectroscopic analyses show that XyNA and dXyNA form stable hairpins. The dXyNA hairpin structure determined by NMR spectroscopy contains a flexible loop that locks the stem into a stable ladder-like duplex with marginal right-handed helicity. The reduced flexibility of the dXyNA duplex observed in the stem of the hairpin demonstrates that folding of dXyNA yields more stable structures described so far.
- Medicinal Chemistry, KU Leuven, Rega Institute for Medical Research, Herestraat 49, Box 1041, 3000, Leuven, Belgium.
Organizational Affiliation: