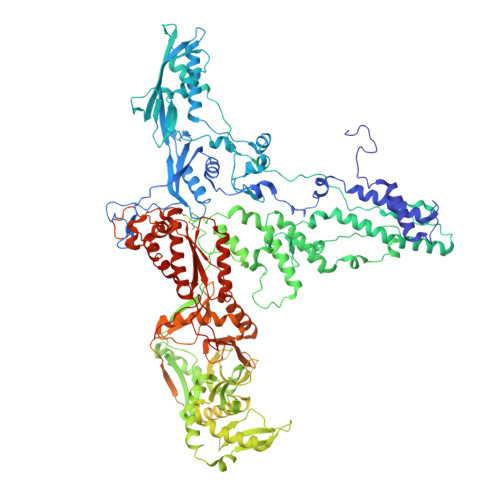
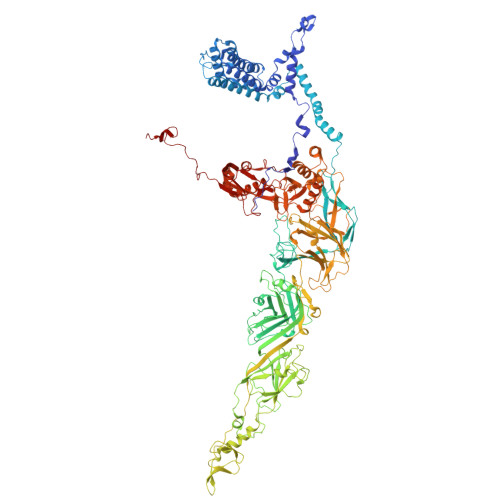
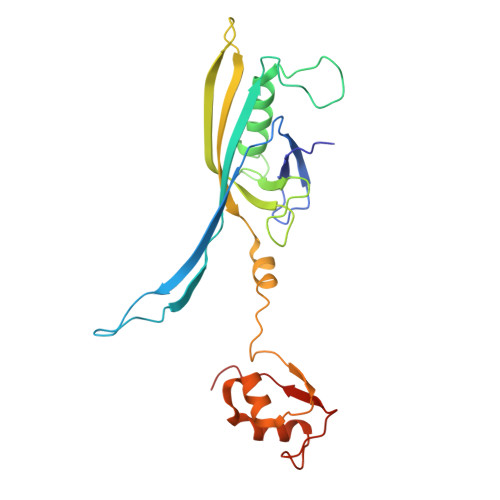
Structure of a thylakoid-anchored contractile injection system in multicellular cyanobacteria.
Weiss, G.L., Eisenstein, F., Kieninger, A.K., Xu, J., Minas, H.A., Gerber, M., Feldmuller, M., Maldener, I., Forchhammer, K., Pilhofer, M.(2022) Nat Microbiol 7: 386-396
- PubMed: 35165386 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41564-021-01055-y
- Primary Citation Related Structures:
7B5H, 7B5I - PubMed Abstract:
Contractile injection systems (CISs) mediate cell-cell interactions by phage tail-like structures, using two distinct modes of action: extracellular CISs are released into the medium, while type 6 secretion systems (T6SSs) are attached to the cytoplasmic membrane and function upon cell-cell contact. Here, we characterized a CIS in the multicellular cyanobacterium Anabaena, with features distinct from extracellular CISs and T6SSs. Cryo-electron tomography of focused ion beam-milled cells revealed that CISs were anchored in thylakoid membrane stacks, facing the cell periphery. Single particle cryo-electron microscopy showed that this unique in situ localization was mediated by extensions of tail fibre and baseplate components. On stress, cyanobacteria induced the formation of ghost cells, presenting thylakoid-anchored CISs to the environment. Functional assays suggest that these CISs may mediate ghost cell formation and/or interactions of ghost cells with other organisms. Collectively, these data provide a framework for understanding the evolutionary re-engineering of CISs and potential roles of these CISs in cyanobacterial programmed cell death.
- Department of Biology, Institute of Molecular Biology & Biophysics, Eidgenössische Technische Hochschule Zürich, Zurich, Switzerland.
Organizational Affiliation: