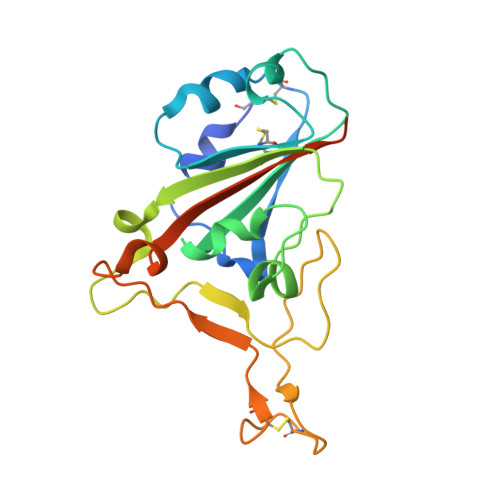
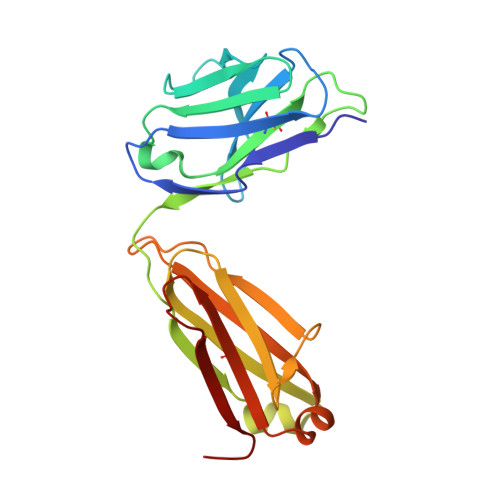
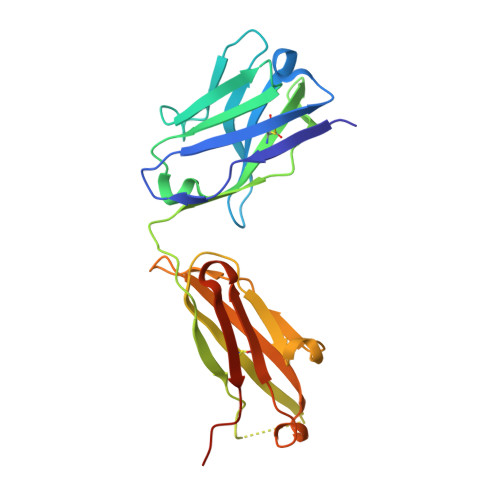
A SARS-CoV-2 neutralizing antibody selected from COVID-19 patients binds to the ACE2-RBD interface and is tolerant to most known RBD mutations.
Bertoglio, F., Fuhner, V., Ruschig, M., Heine, P.A., Abassi, L., Klunemann, T., Rand, U., Meier, D., Langreder, N., Steinke, S., Ballmann, R., Schneider, K.T., Roth, K.D.R., Kuhn, P., Riese, P., Schackermann, D., Korn, J., Koch, A., Chaudhry, M.Z., Eschke, K., Kim, Y., Zock-Emmenthal, S., Becker, M., Scholz, M., Moreira, G.M.S.G., Wenzel, E.V., Russo, G., Garritsen, H.S.P., Casu, S., Gerstner, A., Roth, G., Adler, J., Trimpert, J., Hermann, A., Schirrmann, T., Dubel, S., Frenzel, A., Van den Heuvel, J., Cicin-Sain, L., Schubert, M., Hust, M.(2021) Cell Rep 36: 109433-109433
- PubMed: 34273271 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2021.109433
- Primary Citation Related Structures:
7B3O - PubMed Abstract:
The novel betacoronavirus severe acute respiratory syndrome-coronavirus-2 (SARS-CoV-2) causes a form of severe pneumonia disease called coronavirus disease 2019 (COVID-19). To develop human neutralizing anti-SARS-CoV-2 antibodies, antibody gene libraries from convalescent COVID-19 patients were constructed and recombinant antibody fragments (scFv) against the receptor-binding domain (RBD) of the spike protein were selected by phage display. The antibody STE90-C11 shows a subnanometer IC 50 in a plaque-based live SARS-CoV-2 neutralization assay. The in vivo efficacy of the antibody is demonstrated in the Syrian hamster and in the human angiotensin-converting enzyme 2 (hACE2) mice model. The crystal structure of STE90-C11 Fab in complex with SARS-CoV-2-RBD is solved at 2.0 Å resolution showing that the antibody binds at the same region as ACE2 to RBD. The binding and inhibition of STE90-C11 is not blocked by many known emerging RBD mutations. STE90-C11-derived human IgG1 with FcγR-silenced Fc (COR-101) is undergoing Phase Ib/II clinical trials for the treatment of moderate to severe COVID-19.
- Technische Universität Braunschweig, Institut für Biochemie, Biotechnologie und Bioinformatik, Abteilung Biotechnologie, Spielmannstr. 7, 38106 Braunschweig, Germany.
Organizational Affiliation: