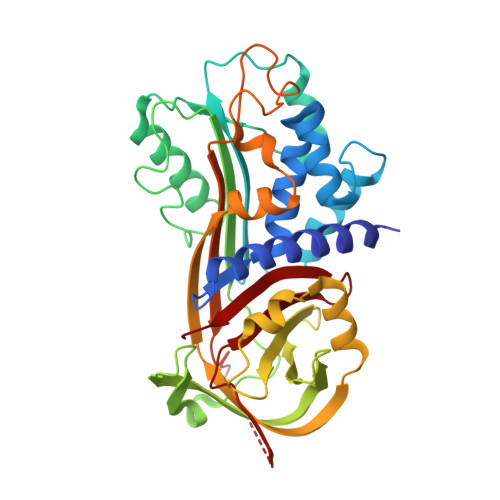
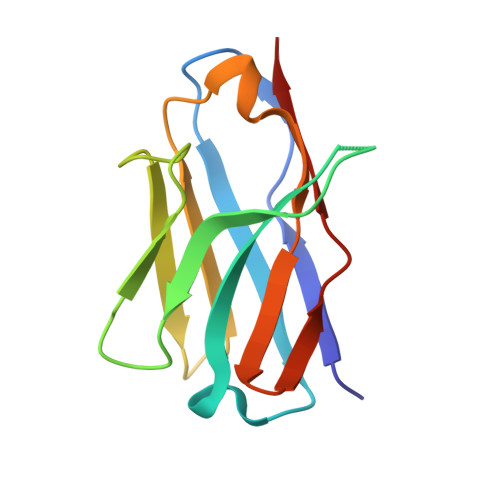
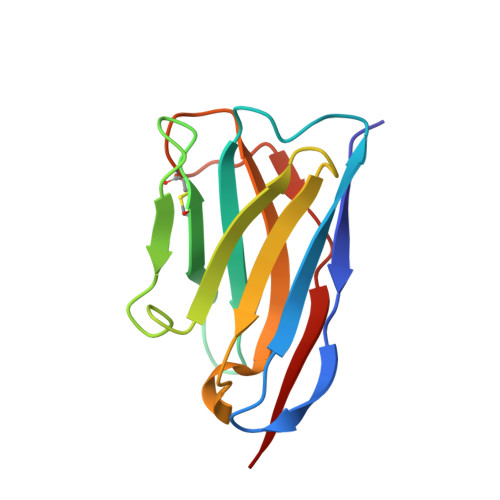
Structural Insight into the Two-Step Mechanism of PAI-1 Inhibition by Small Molecule TM5484.
Sillen, M., Miyata, T., Vaughan, D.E., Strelkov, S.V., Declerck, P.J.(2021) Int J Mol Sci 22
- PubMed: 33540702 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/ijms22031482
- Primary Citation Related Structures:
7AQF, 7AQG - PubMed Abstract:
Plasminogen activator inhibitor-1 (PAI-1), a key regulator of the fibrinolytic system, is the main physiological inhibitor of plasminogen activators. By interacting with matrix components, including vitronectin (Vn), PAI-1 plays a regulatory role in tissue remodeling, cell migration, and intracellular signaling. Emerging evidence points to a role for PAI-1 in various pathological conditions, including cardiovascular diseases, cancer, and fibrosis. Targeting PAI-1 is therefore a promising therapeutic strategy in PAI-1-related pathologies. A class of small molecule inhibitors including TM5441 and TM5484, designed to bind the cleft in the central β-sheet A of PAI-1, showed to be potent PAI-1 inhibitors in vivo. However, their binding site has not yet been confirmed. Here, we report two X-ray crystallographic structures of PAI-1 in complex with TM5484. The structures revealed a binding site at the flexible joint region, which is distinct from the presumed binding site. Based on the structural analysis and biochemical data we propose a mechanism for the observed dose-dependent two-step mechanism of PAI-1 inhibition. By binding to the flexible joint region in PAI-1, TM5484 might restrict the structural flexibility of this region, thereby inducing a substrate form of PAI-1 followed by a conversion to an inert form.
- Laboratory for Therapeutic and Diagnostic Antibodies, Department of Pharmaceutical and Pharmacological Sciences, KU Leuven, B-3000 Leuven, Belgium.
Organizational Affiliation: