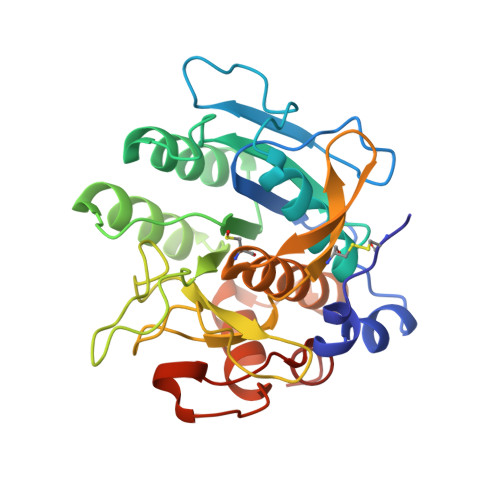
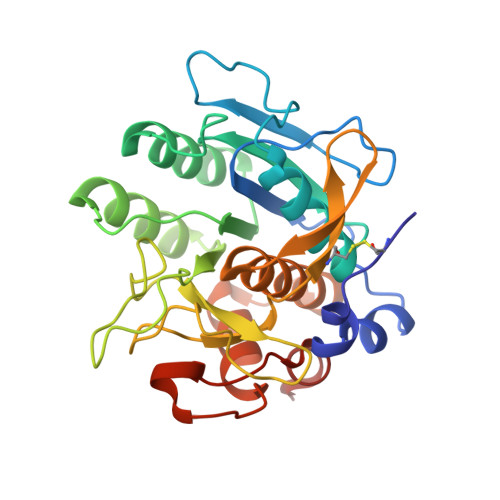
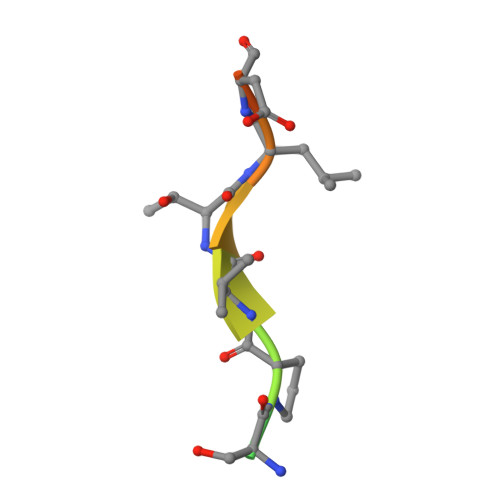
From thiol-subtilisin to omniligase: Design and structure of a broadly applicable peptide ligase.
Toplak, A., Teixeira de Oliveira, E.F., Schmidt, M., Rozeboom, H.J., Wijma, H.J., Meekels, L.K.M., de Visser, R., Janssen, D.B., Nuijens, T.(2021) Comput Struct Biotechnol J 19: 1277-1287
- PubMed: 33717424 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.csbj.2021.02.002
- Primary Citation Related Structures:
7AM3, 7AM4, 7AM5, 7AM6, 7AM7, 7AM8 - PubMed Abstract:
Omniligase-1 is a broadly applicable enzyme for peptide bond formation between an activated acyl donor peptide and a non-protected acyl acceptor peptide. The enzyme is derived from an earlier subtilisin variant called peptiligase by several rounds of protein engineering aimed at increasing synthetic yields and substrate range. To examine the contribution of individual mutations on S/H ratio and substrate scope in peptide synthesis, we selected peptiligase variant M222P/L217H as a starting enzyme and introduced successive mutations. Mutation A225N in the S1' pocket and F189W of the S2' pocket increased the synthesis to hydrolysis (S/H) ratio and overall coupling efficiency, whereas the I107V mutation was added to S4 pocket to increase the reaction rate. The final omniligase variants appeared to have a very broad substrate range, coupling more than 250 peptides in a 400-member library of acyl acceptors, as indicated by a high-throughput FRET assay. Crystal structures and computational modelling could rationalize the exceptional properties of omniligase-1 in peptide synthesis.
- EnzyPep B.V., Brightlands Campus Urmonderbaan 22, 6167 RD Geleen, The Netherlands.
Organizational Affiliation: