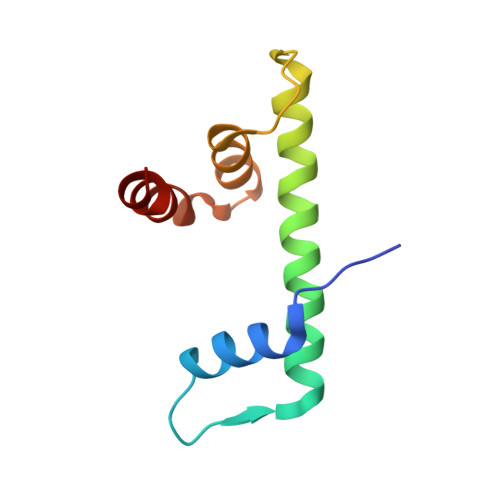
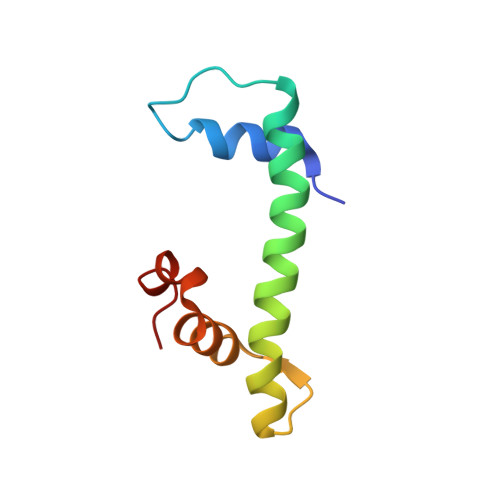
Structural Basis of Inhibition of the Pioneer Transcription Factor NF-Y by Suramin.
Nardone, V., Chaves-Sanjuan, A., Lapi, M., Airoldi, C., Saponaro, A., Pasqualato, S., Dolfini, D., Camilloni, C., Bernardini, A., Gnesutta, N., Mantovani, R., Nardini, M.(2020) Cells 9
- PubMed: 33138093 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/cells9112370
- Primary Citation Related Structures:
7AH8 - PubMed Abstract:
NF-Y is a transcription factor (TF) comprising three subunits (NF-YA, NF-YB, NF-YC) that binds with high specificity to the CCAAT sequence, a widespread regulatory element in gene promoters of prosurvival, cell-cycle-promoting, and metabolic genes. Tumor cells undergo "metabolic rewiring" through overexpression of genes involved in such pathways, many of which are under NF-Y control. In addition, NF-YA appears to be overexpressed in many tumor types. Thus, limiting NF-Y activity may represent a desirable anti-cancer strategy, which is an ongoing field of research. With virtual-screening docking simulations on a library of pharmacologically active compounds, we identified suramin as a potential NF-Y inhibitor. We focused on suramin given its high water-solubility that is an important factor for in vitro testing, since NF-Y is sensitive to DMSO. By electrophoretic mobility shift assays (EMSA), isothermal titration calorimetry (ITC), STD NMR, X-ray crystallography, and molecular dynamics (MD) simulations, we showed that suramin binds to the histone fold domains (HFDs) of NF-Y, preventing DNA-binding. Our analyses, provide atomic-level detail on the interaction between suramin and NF-Y and reveal a region of the protein, nearby the suramin-binding site and poorly conserved in other HFD-containing TFs, that may represent a promising starting point for rational design of more specific and potent inhibitors with potential therapeutic applications.
- Department of Biosciences, University of Milano, Via Celoria 26, 20133 Milano, Italy.
Organizational Affiliation: