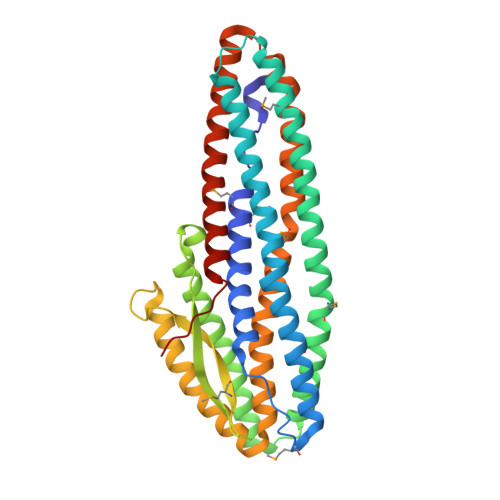
Characterisation of a tripartite alpha-pore forming toxin from Serratia marcescens
Churchill-Angus, A.M., Schofield, T.H.B., Marlow, T.R., Sedelnikova, S.E., Wilson, J.S., Rafferty, J.B., Baker, P.J.(2021) Sci Rep 11: 6447
- PubMed: 33742033 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-021-85726-0
- Primary Citation Related Structures:
6ZZ5, 6ZZH, 7A0G, 7A26, 7A27 - PubMed Abstract:
Tripartite members of the ClyA family of α-PFTs have recently been identified in a number of pathogenic Gram-negative bacteria, including the human pathogen Serratia marcescens. Structures of a Gram-negative A component and a tripartite α-PFT complete pore are unknown and a mechanism for pore formation is still uncertain. Here we characterise the tripartite SmhABC toxin from S. marcescens and propose a mechanism of pore assembly. We present the structure of soluble SmhA, as well as the soluble and pore forms of SmhB. We show that the β-tongue soluble structure is well conserved in the family and propose two conserved latches between the head and tail domains that are broken on the soluble to pore conformational change. Using the structures of individual components, sequence analysis and docking predictions we illustrate how the A, B and C protomers would assemble on the membrane to produce a complete tripartite α-PFT pore.
- Department of Molecular Biology and Biotechnology, University of Sheffield, Firth Court, Western Bank, Sheffield, S10 2TN, South Yorkshire, UK.
Organizational Affiliation: