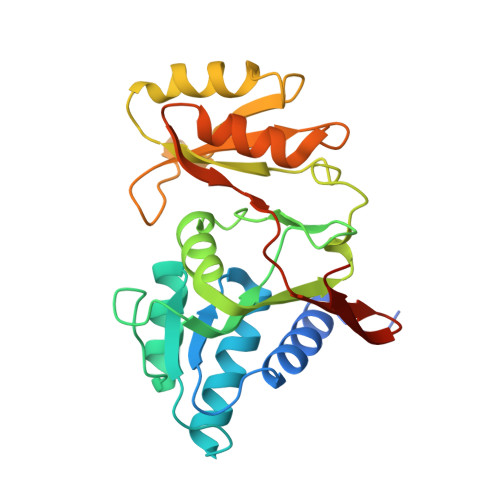
High-Resolution Crystal Structure of Chloroplastic Ribose-5-Phosphate Isomerase from Chlamydomonas reinhardtii -An Enzyme Involved in the Photosynthetic Calvin-Benson Cycle.
Le Moigne, T., Crozet, P., Lemaire, S.D., Henri, J.(2020) Int J Mol Sci 21
- PubMed: 33096784 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/ijms21207787
- Primary Citation Related Structures:
6ZXT - PubMed Abstract:
The Calvin-Benson cycle is the key metabolic pathway of photosynthesis responsible for carbon fixation and relies on eleven conserved enzymes. Ribose-5-phosphate isomerase (RPI) isomerizes ribose-5-phosphate into ribulose-5-phosphate and contributes to the regeneration of the Rubisco substrate. Plant RPI is the target of diverse post-translational modifications including phosphorylation and thiol-based modifications to presumably adjust its activity to the photosynthetic electron flow. Here, we describe the first experimental structure of a photosynthetic RPI at 1.4 Å resolution. Our structure confirms the composition of the catalytic pocket of the enzyme. We describe the homo-dimeric state of the protein that we observed in the crystal and in solution. We also map the positions of previously reported post-translational modifications and propose mechanisms by which they may impact the catalytic parameters. The structural data will inform the biochemical modeling of photosynthesis.
- Laboratoire de Biologie Moléculaire et Cellulaire des Eucaryotes, UMR8226, Institut de Biologie Physico-Chimique, Sorbonne Université, CNRS, 75005 Paris, France.
Organizational Affiliation: