Metal-Mediated Protein-Cucurbituril Crystalline Architectures
Guagnini, F., Engilberge, S., Flood, R.J., Ramberg, K.O., Crowley, P.B.(2020) Cryst Growth Des
Experimental Data Snapshot
Starting Model: experimental
View more details
(2020) Cryst Growth Des
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Fucose-binding lectin protein | 90 | Ralstonia solanacearum | Mutation(s): 1 Gene Names: E7Z57_08365, RSP795_21825, RSP822_19650, RUN39_v1_50103 | 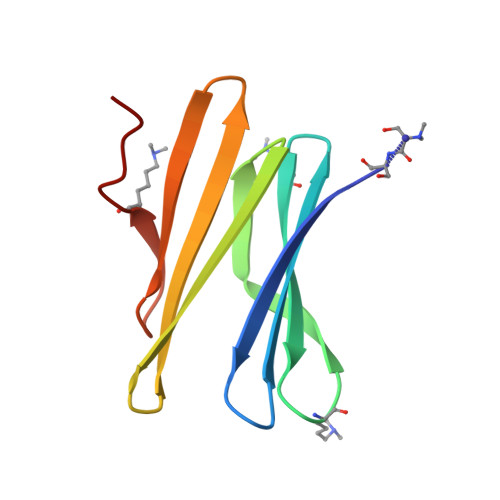 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A0A0S4TLR1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 5 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| QQ7 (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | G [auth A] | cucurbit[7]uril C42 H42 N28 O14 ZDOBFUIMGBWEAB-XGFHMVPTSA-N |  | ||
| BDF Download:Ideal Coordinates CCD File | B [auth A], C [auth A] | beta-D-fructopyranose C6 H12 O6 LKDRXBCSQODPBY-ARQDHWQXSA-N |  | ||
| ZN (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | E [auth A], F [auth A] | ZINC ION Zn PTFCDOFLOPIGGS-UHFFFAOYSA-N |  | ||
| ACT Download:Ideal Coordinates CCD File | D [auth A] | ACETATE ION C2 H3 O2 QTBSBXVTEAMEQO-UHFFFAOYSA-M |  | ||
| NA Download:Ideal Coordinates CCD File | H [auth A], I [auth A] | SODIUM ION Na FKNQFGJONOIPTF-UHFFFAOYSA-N |  | ||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| MLY Query on MLY | A | L-PEPTIDE LINKING | C8 H18 N2 O2 |  | LYS |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 51.057 | α = 90 |
| b = 51.057 | β = 90 |
| c = 102.477 | γ = 120 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| PDB_EXTRACT | data extraction |
| XDS | data reduction |
| Aimless | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Science Foundation Ireland | Ireland | 13/CDA/2168 |
| Science Foundation Ireland | Ireland | 12/RC/2275_P2 |
| Irish Research Council | Ireland | GOIPD/2019/513 |