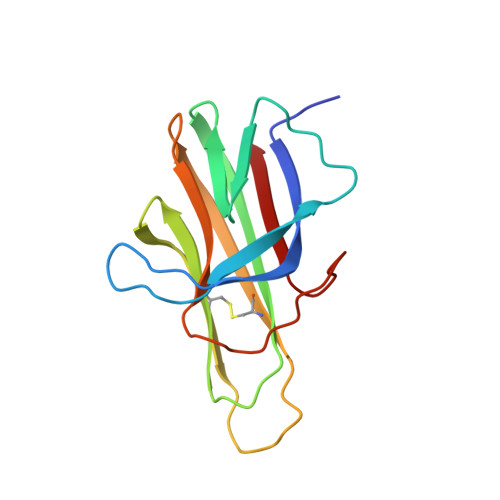
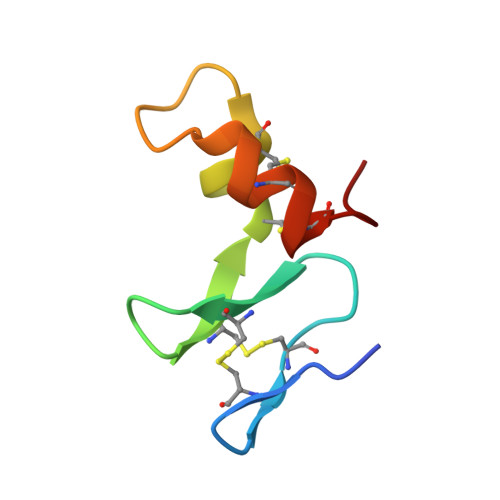
Drosophila TNFRs Grindelwald and Wengen bind Eiger with different affinities and promote distinct cellular functions.
Palmerini, V., Monzani, S., Laurichesse, Q., Loudhaief, R., Mari, S., Cecatiello, V., Olieric, V., Pasqualato, S., Colombani, J., Andersen, D.S., Mapelli, M.(2021) Nat Commun 12: 2070-2070
- PubMed: 33824334 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-22080-9
- Primary Citation Related Structures:
6ZSY, 6ZSZ, 6ZT0 - PubMed Abstract:
The Drosophila tumour necrosis factor (TNF) ligand-receptor system consists of a unique ligand, Eiger (Egr), and two receptors, Grindelwald (Grnd) and Wengen (Wgn), and therefore provides a simple system for exploring the interplay between ligand and receptors, and the requirement for Grnd and Wgn in TNF/Egr-mediated processes. Here, we report the crystallographic structure of the extracellular domain (ECD) of Grnd in complex with Egr, a high-affinity hetero-hexameric assembly reminiscent of human TNF:TNFR complexes. We show that ectopic expression of Egr results in internalisation of Egr:Grnd complexes in vesicles, a step preceding and strictly required for Egr-induced apoptosis. We further demonstrate that Wgn binds Egr with much reduced affinity and is localised in intracellular vesicles that are distinct from those containing Egr:Grnd complexes. Altogether, our data provide insight into ligand-mediated activation of Grnd and suggest that distinct affinities of TNF ligands for their receptors promote different and non-redundant cellular functions.
- IEO, European Institute of Oncology IRCCS, Milan, Italy.
Organizational Affiliation: