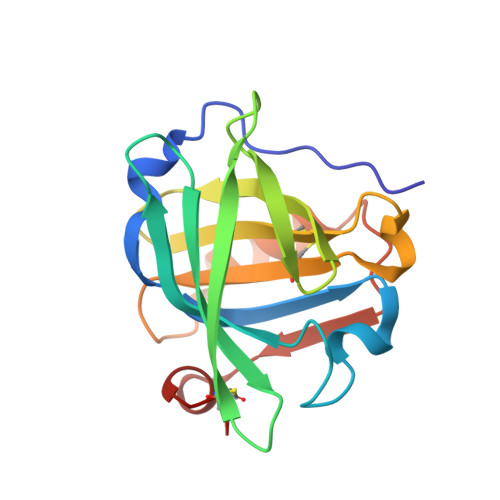
Cisplatin binding to beta-lactoglobulin: a structural study.
Balasco, N., Ferraro, G., Loreto, D., Iacobucci, I., Monti, M., Merlino, A.(2020) Dalton Trans 49: 12450-12457
- PubMed: 32852026 Search on PubMed
- DOI: https://doi.org/10.1039/d0dt02582h
- Primary Citation Related Structures:
6ZSQ, 6ZSR - PubMed Abstract:
β-Lactoglobulin is a major globular milk whey carrier with potential applications as an oral drug delivery system. Herein, the interactions between β-lactoglobulin and cisplatin are investigated by UV-Vis absorption spectroscopy, circular dichroism, X-ray crystallography and electrospray ionization mass spectrometry. Structural data indicate that the protein retains its conformation upon cisplatin binding. Pt-containing fragments bind the side chains of Met7, His146 and Lys8, with the number of binding sites increasing over time. Mass spectrometry data indicate that [Pt(NH3)2Cl+], [Pt(NH3)2OH22+] and [Pt(NH3)22+] fragments interact with β-lactoglobulin; up to 3 cisplatin fragments can bind the protein and the number of cisplatin binding sites increases over time. This work opens a new pathway in pharmaceutical studies based on a rational design of metal-based drug/β-lactoglobulin adducts as delivering vehicles of metallodrugs.
- Institute of Biostructures and Bioimaging, CNR, Via Mezzocannone 16, 80134 Napoli, Italy.
Organizational Affiliation: