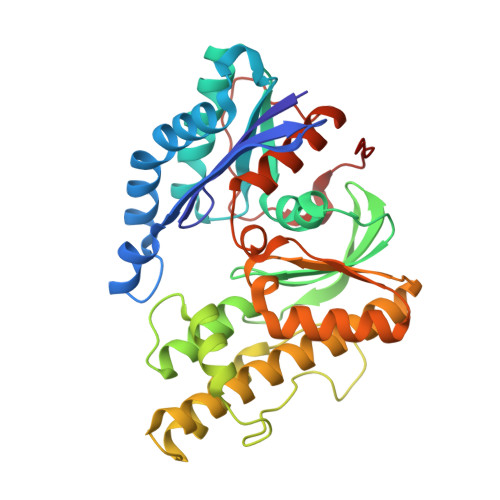
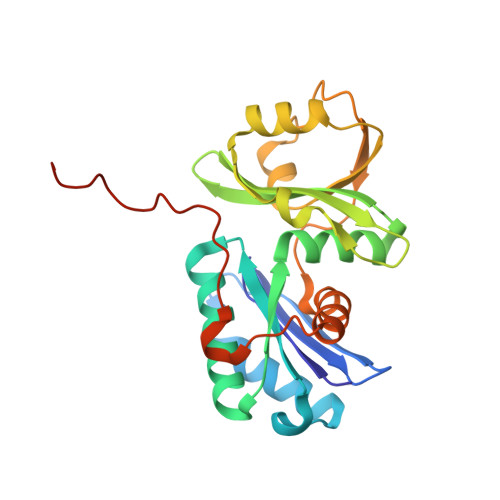
Structure of a reaction intermediate mimic in t6A biosynthesis bound in the active site of the TsaBD heterodimer from Escherichia coli.
Kopina, B.J., Missoury, S., Collinet, B., Fulton, M.G., Cirio, C., van Tilbeurgh, H., Lauhon, C.T.(2021) Nucleic Acids Res 49: 2141-2160
- PubMed: 33524148 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkab026
- Primary Citation Related Structures:
6Z81 - PubMed Abstract:
The tRNA modification N6-threonylcarbamoyladenosine (t6A) is universally conserved in all organisms. In bacteria, the biosynthesis of t6A requires four proteins (TsaBCDE) that catalyze the formation of t6A via the unstable intermediate l-threonylcarbamoyl-adenylate (TC-AMP). While the formation and stability of this intermediate has been studied in detail, the mechanism of its transfer to A37 in tRNA is poorly understood. To investigate this step, the structure of the TsaBD heterodimer from Escherichia coli has been solved bound to a stable phosphonate isosteric mimic of TC-AMP. The phosphonate inhibits t6A synthesis in vitro with an IC50 value of 1.3 μM in the presence of millimolar ATP and L-threonine. The inhibitor binds to TsaBD by coordination to the active site Zn atom via an oxygen atom from both the phosphonate and the carboxylate moieties. The bound conformation of the inhibitor suggests that the catalysis exploits a putative oxyanion hole created by a conserved active site loop of TsaD and that the metal essentially serves as a binding scaffold for the intermediate. The phosphonate bound crystal structure should be useful for the rational design of potent, drug-like small molecule inhibitors as mechanistic probes or potentially novel antibiotics.
- Pharmaceutical Sciences Division, School of Pharmacy, University of Wisconsin, Madison, WI 53705, USA.
Organizational Affiliation: